AbstractPurposeCurrent variability in methods for tumor mutational burden (TMB) estimation and reporting demonstrates the urgent need for a homogeneous TMB assessment approach. Here, we compared TMB distributions in different cancer types using two customized targeted panels commonly used in clinical practice.
Materials and MethodsTMB spectra of 295- and 1021-gene panels in multiple cancer types were compared using targeted next-generation sequencing (NGS). The TMB distributions across a diverse cohort of 2,332 cancer cases were then investigated for their associations with clinical features. Treatment response data were collected for 222 patients who received immune-checkpoint inhibitors (ICIs) and their homologous recombination DNA damage repair (HR-DDR) and programmed death-ligand 1 (PD-L1) expression were additionally assessed and compared with the TMB and response rate.
ResultsThe median TMB between gene panels was similar despite a wide range in TMB values. The highest TMB was eight and 10 in patients with squamous cell carcinoma and esophageal carcinoma according to the classification of histopathology and cancer types, respectively. Twenty-three out of 103 patients (22.3%) were HR-DDR–positive and could benefit from ICI therapy; out of those 23 patients, seven patients had high TMB (p=0.004). Additionally, PD-L1 expression was not associated with TMB or treatment response among patients receiving ICIs.
IntroductionThe immune system plays a pivotal role not only in cancer recognition but also in monitoring cells for neoantigen expression and modulating antitumor activity [1]. Recently, the use of immune checkpoint inhibitors (ICIs), such as cytotoxic T lymphocyte-associated antigen 4 [2,3], programmed cell death receptor-1 (PD-1) and its ligand (PD-L1) [4,5], has shown remarkable clinical benefits in the treatment of melanoma, lung, renal cell, and prostate cancers [6–8]. However, since only a subset of patients respond to ICIs [9], it is of paramount importance to identify biomarkers that predict treatment response and outcomes, allowing more efficient and timely treatment.
Defective DNA mismatch repair and PD-L1 expression are the only two predictive biomarkers for immunotherapy approved by the Food and Drug Administration [10]. Several studies have demonstrated that the presence of these biomarkers is associated with greater numbers of somatic mutations and tumor neoantigens, influencing the sensitivity of different tumor types (e.g., melanoma, non-small cell lung cancer, and mismatch-repair deficient tumors) to ICIs [11]. Tumor mutational burden (TMB), an indirect measure of tumor-derived neoantigens, is measured through whole-exome sequencing (WES) or cancer gene panels and is another extensively studied biomarker that shows promising potential in predicting treatment response to ICIs [12,13]. The homologous recombination DNA damage repair (HR-DDR) pathway mutations have also been identified as potential indicators of the treatment response to anticancer therapies [14]. To date, no research has been conducted to assess the correlation between HR-DDR status and TMB or whether these biomarkers could jointly better stratify patients regarding their predicted ICI response.
To better understand the TMB across the spectrum of human cancers, we performed a cohort study of 2,332 cancer patients who underwent 295- and 1021-gene panel genetic testing using a next-generation sequencing (NGS) platform at the Sun Yat-sen University Cancer Center (SYSUCC) in Guangzhou, China. This study aimed to characterize TMB in detail across various cancer types and understand the significance of combining TMB and HR-DDR status in predicting ICI treatment response using PD-L1 expression as the reference.
Materials and Methods1. Patient selectionData from 2,332 patients who underwent genomic profiling with hybridization capture-based NGS assay from January 1, 2017, to January 31, 2020 at the SYSUCC were retrospectively retrieved. Eligible patients were defined as those with a pathologically confirmed cancer diagnosis. We collected clinical data concerning age, sex, smoking status, percentage of PD-L1 membranous staining of tumor cells, cancer stage, and family history for all patients.
A subset of patients (n=222) received immunotherapy and their treatment response was characterized as complete or partial response (CR/PR), stable disease (SD), or progressive disease based on the Response Evaluation Criteria in Solid Tumor ver. 1.1 criteria (ref). We defined the effectiveness of ICI treatment as durable clinical benefit (DCB) or no durable benefit (NDB). DCB was defined as CR/PR or SD for at least one month, whereas NDB was defined as disease progression within 1 month after the start of ICI treatment. The study protocol is summarized in Fig. 1.
2. Tumor samples and targeted NGSParaffin-embedded tumor biopsy or surgical samples were retrieved from all patients, with a minimum of 20% tumor cells within each tissue for sequencing, which was assessed by the examination of hematoxylin and eosin–stained slides by a pathologist (Y.-F.F.). We used two targeted sequencing assays, including the 295 OncoScreen panel containing whole exons of 287 genes and selected introns of 22 genes (Burning Rock Biotech Ltd., Guangzhou, China) (S1 Table) and the 1021-gene panel containing whole exons and selected introns of 288 genes and selected regions of 733 genes (Geneplus-Beijing, Beijing, China) (S1 Table). These two gene panels are reliable and have a good correlation in evaluating TMB when compared to TMB assessed using WES, respectively (S2 Fig.). Briefly, DNA was extracted from the retrieved tumor samples and matched peripheral blood or adjacent tissue samples and were used for TMB assessment and filtering germline mutations across multiple cancer types. DNA fragmentation was conducted using a Covaris M220 Focused-ultrasonicator (Woburn, MA), followed by end-repair, phosphorylation, and adaptor ligation. Barcoded libraries were generated and sequenced for all exons, and selected introns of a custom panel of 295 (for 1,740 patients) and 1,021 (for 592 patients) genes, respectively. All indexed libraries were sequenced to a minimal unique coverage depth of 100× on a NextSeq 500 platform (Illumina, San Diego, CA) and Gene+Seq-2000 (Geneplus-Beijing Institute, Suzhou, China). Adaptor sequences and low-quality reads were removed, and the clean reads in the FASTQ format were mapped to the reference human genome (hg19) using the Burrows-Wheeler Aligner (ver. 0.7.10-r1039). Local alignment optimization, variant calling, and annotation were performed using GATK (ver. 3.4-46-gbc02625), MuTect, and VarScan, respectively. To normalize the somatic TMB across the 295- and 1021-gene panels, the total number of mutations was divided by the number of coding regions captured in each panel (1.02 and 1.60 mega-bases [Mb], respectively).
3. TMB assessmentThe calculation of TMB was performed using Ion Reporter Analysis Software v5.10 (IR) using the Oncomine Tumor Mutation Load w2.0 workflow (Thermo Fisher Scientific, Waltham, MA). From both the 295- and 1021-gene panel targeted profiling samples, we calculated the numbers of somatic missense mutations, nonsense mutations, and coding indels by the number of exonic bases with at least 500× coverage, and display the findings as the number of mutations per Mb of captured genome. Fusions, copy number variations, and non-coding mutations were not counted [15]. The default limit of detection was set at 5% allelic frequency and adjusted to 10%, depending on the presence of potential deamination artifacts. Subsequently, we examined the frequency of common oncogenic mutations, such as mutations in TP53, KRAS, and PIK3CA, found in cancers and their association with TMB. We also analyzed the mutation status of homologous recombination deficiency (HRD) genes, which are defined as frame-shift mutations, premature stop codons, mutations shown to disrupt natural splicing, and point mutations [16] associated with the TMB spectrum in a subset of 222 patients.
4. Immunohistochemistry for PD-L1 expressionFormalin-fixed, paraffin-embedded tissue blocks were sectioned at 4 μm for PD-L1 expression using monoclonal antibodies against Ventana PD-L1 (SP142 assay on Ventana Benchmark Ultra, Ventana Medical Systems, Tucson, AZ), based on previously described methods [17]. The sections were then dried and adhered to the slides by baking at 60°C for 1 hour. Immunohistochemical staining was conducted on automated platforms according to the manufacturer’s instructions. According to the current convention for PD-L1 expression assessment, the percentage of tumor cells with membranous staining was scored using the Olympus micro-scope by two pathologists (Y.-F.F. and Y.H.). In cases of disagreement, a consensus was reached after joint review using a multihead microscope. Different thresholds for PD-L1 expression were determined to be ≤ 1% and > 1%.
5. Statistical analysisDescriptive statistics were used to characterize the demographic and clinical features of patients using the mean or median values for continuous variables. The significance of differences in baseline characteristics was assessed using the unpaired t test and Fisher exact test among patients tested using the two gene panels. The nonparametric Mann-Whitney U test was used to test for differences in the median values of TMB between the 295- and 1021-gene panels. The correlations between TMB, HR-DDR status, and ICI treatment response were examined using Spearman rank correlation coefficients. All statistical analyses were performed using Stata ver. 14.0 (StataCorp LLC, College Station, TX), GraphPad Prism 7.04 (GraphPad Software Inc., San Diego, CA), and R ver. 3.3.3 software (http://www.R-project.org). A p-value < 0.05 was considered statistically significant.
Results1. Patient characteristicsThe demographic and clinical characteristics of the enrolled patients are detailed in Table 1. The median age at diagnosis was 55 years (range, 1 to 92 years); 53.7% of patients were male and 16.4% were current or former smokers. Smoking status was known for 2,103 patients, including 383 smokers (16.4%). The distribution of cancer types is illustrated in Fig. 2A. The most common cancer type was colorectal cancer (CRC; n=681, 29.2%), followed by lung cancer (n=510, 21.9%), melanoma (n=232, 10.0%), and gastric cancer (n=143, 6.3%). Histologically, 1,582 of the cases (67.8%) were adenocarcinoma, irrespective of tumor origin (Fig. 2B). The other histopathological types included mesenchymal tumors, squamous cell carcinoma, and adeno-squamous cell carcinoma.
2. Landscape of TMBIn total, 2,332 tumor samples were successfully sequenced using the 295- and 1021-gene panels. The overall median TMB was 6 (range, 2 to 227) and 7 (range, 2 to 802) mutations per Mb for the 295- and 1021-gene panels, respectively (p < 0.001) (S3 Fig.). This correlation of the TMB value in the 295- and 1021-gene panel was similar (R2=0.9655) (S4 Fig.). Not surprisingly, sex and smoking status were associated with median TMB (p < 0.001 for sex and smoking status) (S5 Fig.).
Among all cancer types, the median TMB ranged from 3 to 10 with the lowest median of 3 (range, 2 to 9) noted in salivary gland carcinoma (n=46) and the highest median of 10 (range, 3 to 41) noted in esophageal carcinoma (n=19) (Fig. 3A, S6 Table). The median TMB was 7 (range, 2 to 802) among CRC patients, and was greater when using the 1021-gene panel (median, 8; range, 2 to 802) than when using the 295-gene panel (median, 7; range, 2 to 145) (p < 0.001) (Fig. 3B). Among patients with endometrial, gallbladder, liver, or gastric cancer, the difference in median TMB was also statistically significant when comparing the 295- and the 1021-gene panels (p < 0.05) (Fig. 3B, S6 Table). Although a similar trend was also noted for lung cancer, the difference was not statistically significant (median, 6; range, 2 to 37 for the 295-gene panel; and median, 7; range, 2 to 55 for the 1021-gene panel [p=0.3066]) (Fig. 3B). Comparisons of median TMB values for other cancer types are shown in S6 Table.
The most common histological subtype was adenocarcinoma (67.8%, 1,582/2,332). The median TMB was 6 among patients with adenocarcinoma, and there was a statistically significant difference between the two gene panels (median, 6; range, 2 to 145; median, 7; range, 2 to 802; p < 0.001) (Fig. 3C and D, S7 Table). The median TMB was 10 in squamous cell carcinoma when using the 1021-gene panel, which was higher than that when using the 295-gene panel (p=0.003) (Fig. 3, S7 Table). Relatively low median TMB values were found among patients with mesenchymal tumors (median, 4; range, 2 to 227) and blastoma (median, 4; range, 2 to 12) (Fig. 3C), with no significant difference observed between the two panels (Fig. 3D). Please refer to S7 Table for more details.
In lung cancer, the most common histological subtype was adenocarcinoma (401/507, 79.1%). The median TMB was higher for squamous cell lung cancer than for adenocarcinoma (p=0.014) (S8A Fig.). Regarding CRC, there was no difference in median TMB between the left- and right-sided colon, irrespective of the gene panel used (p=0.129 and p=0.745) (S9A and S9B Fig.).
Mutations in TP53 were observed among 1,241 of the 1,993 patients analyzed (62.2%) (S10A Fig.). KRAS mutations were observed in 517 of the 1,993 patients (26.0%), including 304 patients with CRC (58.8%), indicating that mutated KRAS was probably associated with higher TMB in CRC patients. PIK3CA mutations were enriched in nine out of 14 patients with a median TMB > 100. The relationship of mutation types and frequencies in the other main genes with TMB are summarized in S8A Fig. Mutations in ARID1A were most frequent among patients with high TMB (25.3%, 191/754), followed by BRCA2 (21.4%, 161/754) and ATM (19.0%, 143/754) (S10B Fig.).
3. TMB, HR-DDR status, and immunotherapy responseThe characteristics of the 222 patients treated with ICIs are described in Table 2, mainly comprising patients with melanoma (n=107, 48.2%) and lung cancer (n=34, 15.3%). The lists of HR-DDR–related genes in each panel are included in S11 Table, and the detailed mutations are shown in S12 Table. The prevalence of HR-DDR mutations was 15.8% (n=35), and the proportion of cases with DCB was 46.8% (n=103). We did not observe a significant difference in treatment response between cancer types in the subset of patients receiving ICIs (S13 Table). Among the patients with colorectal cancer, 13 out of 16 patients had a DCB response, partially due to the positive status of the HR-DDR pathway (12/16) and high TMB in patients with TMB > 10 (14/16). However, there were no positive correlations between ICI treatment and HR-DDR–positive status and/or high TMB in patients with melanoma or lung cancer. We found a higher median TMB among patients with HR-DDR mutations than among those without HR-DDR mutations (p < 0.001) (Fig. 4). Interestingly, patients with HR-DDR mutation and high TMB had better disease control than patients with low TMB (p < 0.001) (Fig. 4). Specifically, in the HR-DDR mutant subgroup, there were two CRC patients with Lynch syndrome that had a DCB with a median TMB of 79 and 80 Muts/Mb, respectively. Interestingly, we found that 14 patients exhibited a median TMB value > 100, of whom eight patients had CRC. More importantly, 6 of 14 were patients with microsatellite instability–high (MSI-H) mutations and four patients had POLE/POLD1 mutations. The genomic landscapes of the remaining four patients without MSI-H or POLE/POLD1 mutations are also provided in S14 Fig.
Among the 116 in whom PD-L1 expression could be evaluated (S15 Fig., S16 Table), 52 patients were defined as having PD-L1 expression ≤ 1% and had a median TMB of 5. The remaining 64 patients were classified as having PD-L1 expression > 1% and had a median TMB of 5. There was no statistically significant correlation between TMB and PD-L1 expression. No clear difference was detected in the ICI response by PD-L1 expression status, without taking TMB into account.
DiscussionIn this study, we evaluated and compared TMB distributions using two commercially customized NGS panels in 2,332 cancer patients and further assessed the treatment response in relation to TMB and HRD status in a subset of patients who received ICI therapy (n=222). The novelty of this study includes a comprehensive description of TMB among more than 20 cancer types with detailed information on pathological subtypes. The finding of a better treatment response in relation to high TMB plus HR-DDR mutant status among ICI-treated patients is another novelty.
The targeted panel size has been linked to the accuracy of TMB estimation [15]. A sequencing panel comprising more than 300 cancer-related genes can help predict TMB, whereas a panel comprising fewer than 150 genes has poor performance [18]. However, Ma et al. [19] reported that the 106-CDS (coding sequencings) mutation panel was also reliable in the estimation of TMB. We used both a 1.02-Mb (295 genes) and a 1.60-Mb (1021 genes) NGS panel, which have been used previously in many investigations [14,19–23] and are both sufficient for accurate TMB estimation, in the present study. The median TMB was similar (6 vs. 7 mutations/Mb), indicating that diagnostic NGS panels targeting several hundred genes could accurately measure TMB and might be clinically useful.
The present results show that esophageal carcinoma patients had comparatively higher median TMB than other cancer patients, and most of these patients had squamous cell carcinoma. Smokers had a greater TMB than non-smokers. Many studies have demonstrated that smokers have higher TMB than non-smokers [24] and smoking is a major risk factor for lung squamous cell carcinoma [25]. Accordingly, we observed a higher TMB in patients with squamous cell lung cancer in the present study. Furthermore, a lower TMB was found in patients with salivary gland carcinoma than in patients with other cancers, in agreement with a previous study reporting low TMB (range, 3 to 6) in patients with salivary gland carcinoma [26].
Mutations in a number of genes have been found to be responsible for increased TMB [27], which is important to better understand this key driver of cancer progression and its related molecular mechanisms. Altered microsatellite loci and increased TMB are known to correlate positively [28]. POLE/POLD1 is a key pathway for DNA replication in which defects can lead to an increased somatic mutation rate [29]. In the present study, the highest mutation rate was noted for TP53. This is similar to other studies showing that loss of TP53 following somatic mutation, copy number loss, and epigenetic silencing are very common in cancer and can be associated with increased mutation frequency [30]. Our study showed that KRAS gene mutations mostly occurred in CRC patients who commonly had a high TMB, which was in line with a previous study demonstrating that the median TMB was higher among KRAS-mutant patients than in KRAS-wild patients [31].
In addition to TMB, other molecular features have also been hypothesized to affect the treatment response of ICIs. For example, HR-DDR deficiency has been associated with a better response to platinum-based neoadjuvant therapy in breast cancer [32]. Perturbations of HR-DDR are considered deleterious to genomic integrity [16]. Several studies have reported that PD-1/PD-L1 expression has limited predictive power for treatment response to ICIs, although ICIs targeting PD-1/PD-L1 offer a novel treatment avenue in some cancer types, such as CRC, lung cancer, metastatic urothelial carcinoma, and melanoma [10,33,34]. In a clinical trial, TMB was shown to be more strongly associated with response to ICIs than PD-L1 expression [35]. To date, little has been reported regarding the relationship between alterations of HR-DDR pathways, PD-L1 expression, and TMB. A recent study showed that neither TMB nor PD-L1 expression correlated with the ICI response and TMB was not significantly associated with PD-L1 expression in metastatic renal cell carcinoma. However, an enrichment of mutations in HR-DDR in patients with disease control was notably displayed compared with the progression disease group [36]. In the present study, we described the correlations between HR-DDR status, PD-L1 expression, and TMB. Patients with HR-DDR–positive status had a higher TMB and better treatment response to ICIs than patients with HR-DDR–negative status. There was no significant association between PD-L1 expression and ICI response, indicating that HR-DDR and TMB might be better markers for ICI response than PD-L1 expression. We also found that 11 out of 14 patients with a TMB > 100 had HR-DDR gene mutations, corroborating a recent study that found alterations in DDR genes were strongly associated with a clinical benefit in patients with metastatic urothelial carcinoma [33].
The present study has two limitations. We did not use a TMB cutoff value in analyses mainly because it is a continuous variable without a clearly defined cutoff point below which responses do not occur and above which a response is guaranteed. Each cancer type might have a specific cutoff value that may be identified with the accumulation of more cases of each cancer type in a subsequent study. The current study had a large sample size, whereas a small number of patients who lacked follow-up data were analyzed in the association of ICI response and TMB and HR-DDR status, which may limit the power of conclusions. A large cohort size is needed to determine the causal relationship between HR-DDR status and TMB.
Our findings show that two customized NGS assays targeting 1.02- and 1.6-Mb of the coding genome could accurately assess TMB in clinical settings, and patients with HR-DDR gene alterations are more likely to have higher TMB and experience better responses to ICIs. Additional investigation is warranted to evaluate the mechanisms that link together HR-DDR alterations, TMB, and responses to immunotherapy, which might represent a useful predictive biomarker for ICI therapy.
Electronic Supplementary MaterialSupplementary materials are available at Cancer Research and Treatment website (https://www.e-crt.org).
NotesEthical Statement For the use of the clinical data for research purposes, prior written informed consents from all patients and approval from the Institute Research Ethics Committee of Sun Yat-sen University Cancer Center (B2020-344-01) were obtained. Author Contributions Conceived and designed the analysis: Wang HY, Wang F. Collected the data: Deng L, Li YQ, Zhang X, Long YK, Zhang X, Feng YF, He Y, Yang XH, Wang F. Contributed data or analysis tools: Wang HY, Deng L, Zhang X, Long YK, Feng YF, He Y, Tang T, Yang XH. Performed the analysis: Wang HY. Wrote the paper: Wang HY. AcknowledgmentsThis study was partially supported by Natural Science Foundation of Guangdong Province (2020A1515010313) and the National Natural Science Foundation of China (81602468). We thank Dr. Seeruttun Sharvesh Raj for the professional English language revision of this manuscript.
Fig. 1Flowchart illustrating the analytic workflow of the study comprising of TMB calculations from the 295- and 1021-customized sequencing panels. HR-DDR, homologous recombination DNA damage repair; NGS, next-generation sequencing; TMB, tumor mutational burden. 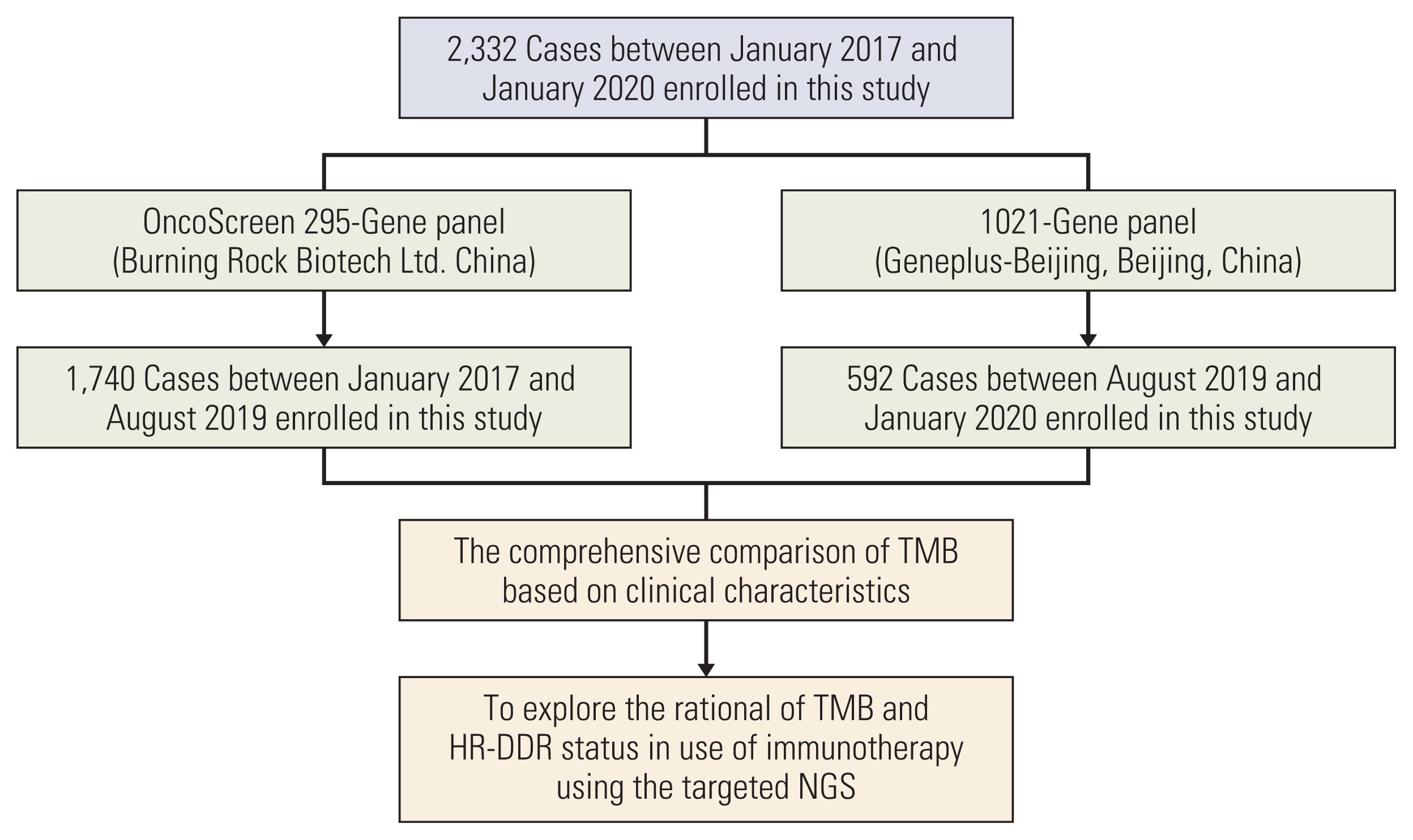
Fig. 2The number of patients was calculated using the 295- and 1021-gene panels, and defined by cancer types (A) and pathological subtypes (B), respectively. SCC, squamous cell carcinoma. Unknown cancer types denote the sites of primary tumor were not available. 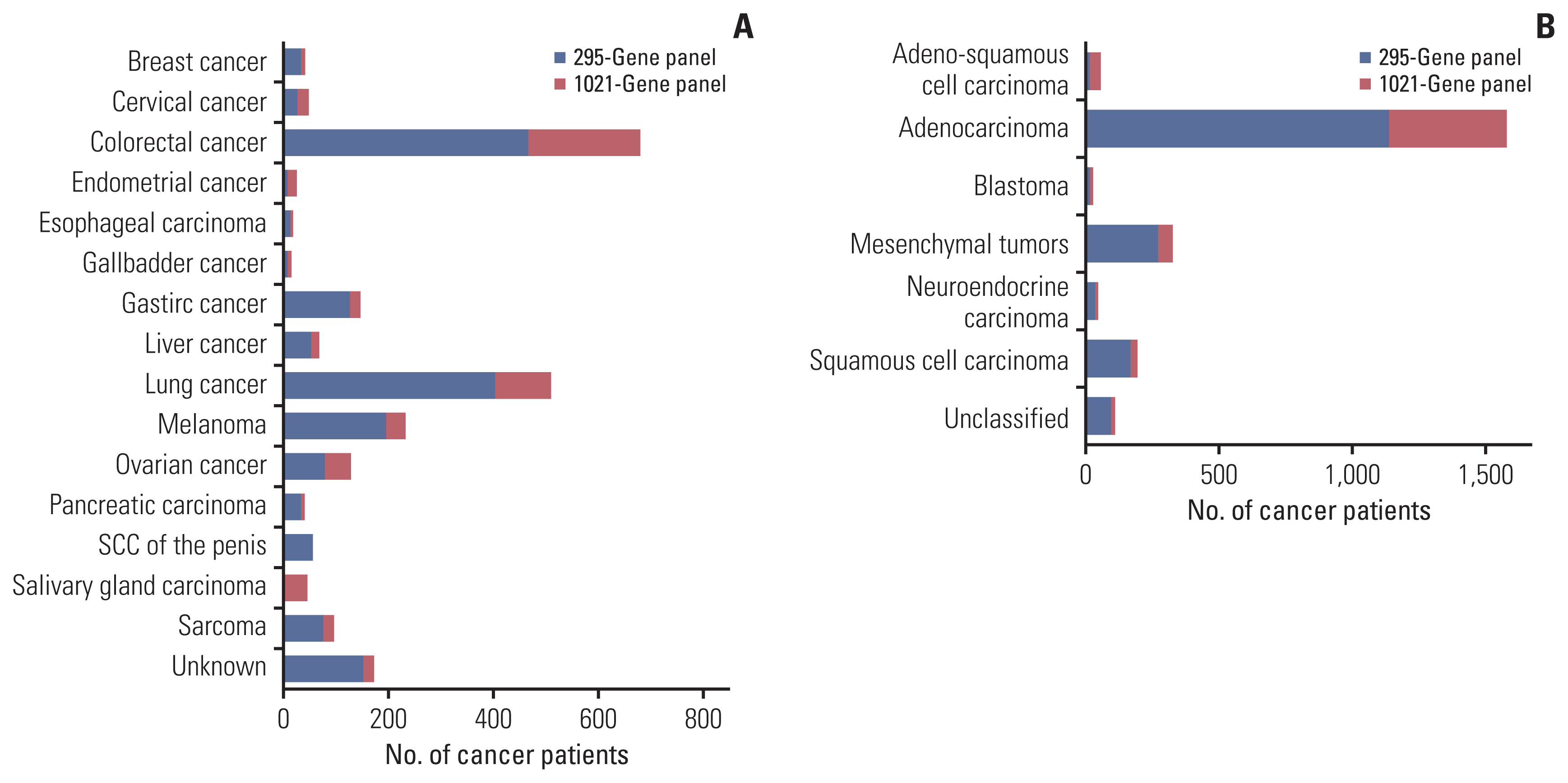
Fig. 3TMB distributions across the different cancer types and pathological subtypes. Overview of the median TMB (log10-transformed) across different cancers (A) and pathological types (C). The comparison of TMB value (log10-transformed) defined by two different gene panels in cancers (B) and pathological types (D). SCC, squamous cell carcinoma; TMB, tumor mutational burden. 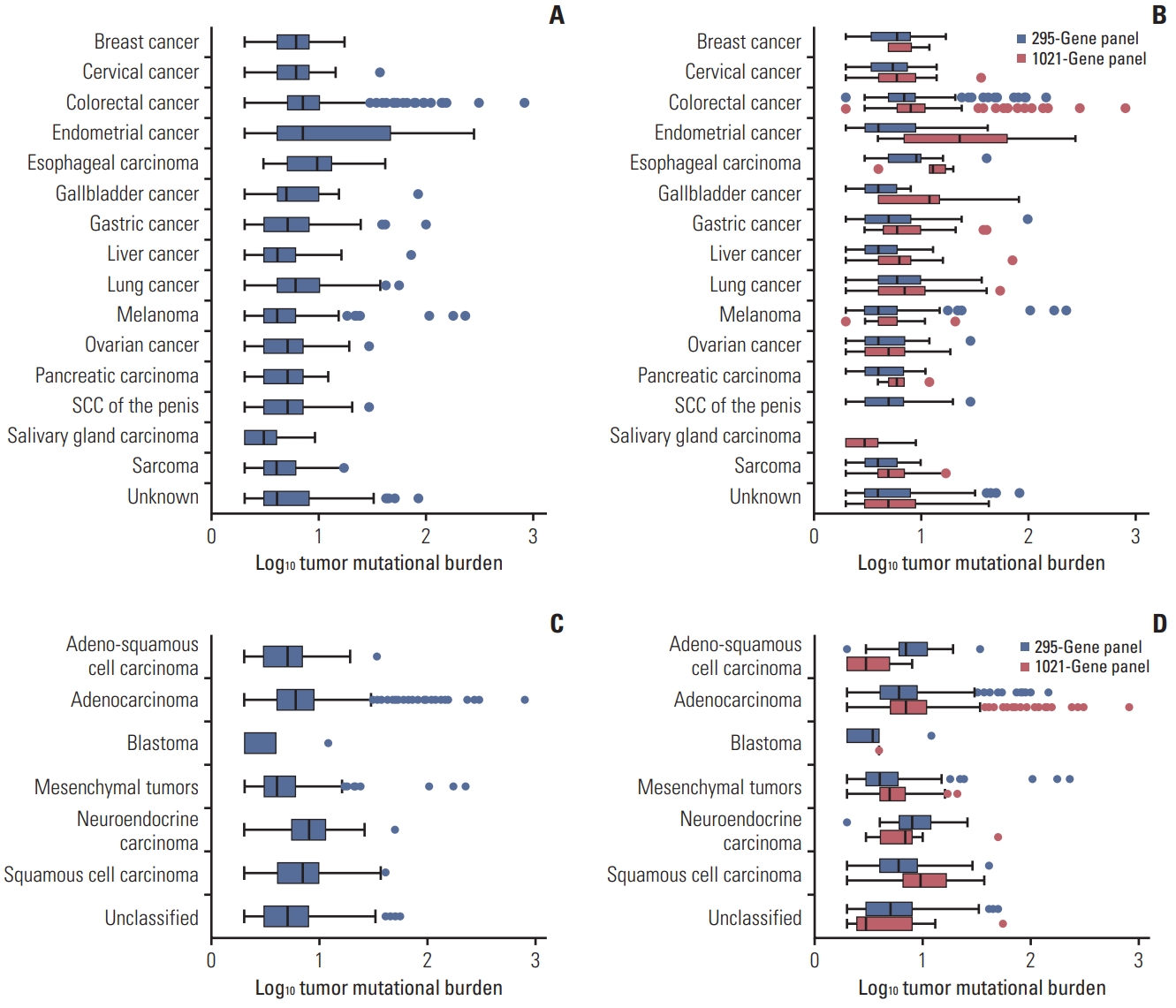
Fig. 4The efficacy associated with TMB, HR-DDR mutation status PD-L1 expression among patients who received ICI treatment. DCB, durable clinical benefit; HR-DDR, homologous recombination DNA damage repair; ICIs, immunotherapy inhibitors; NDB, no durable benefit; PD-L1, programmed death ligand-1; TMB, tumor mutational burden. 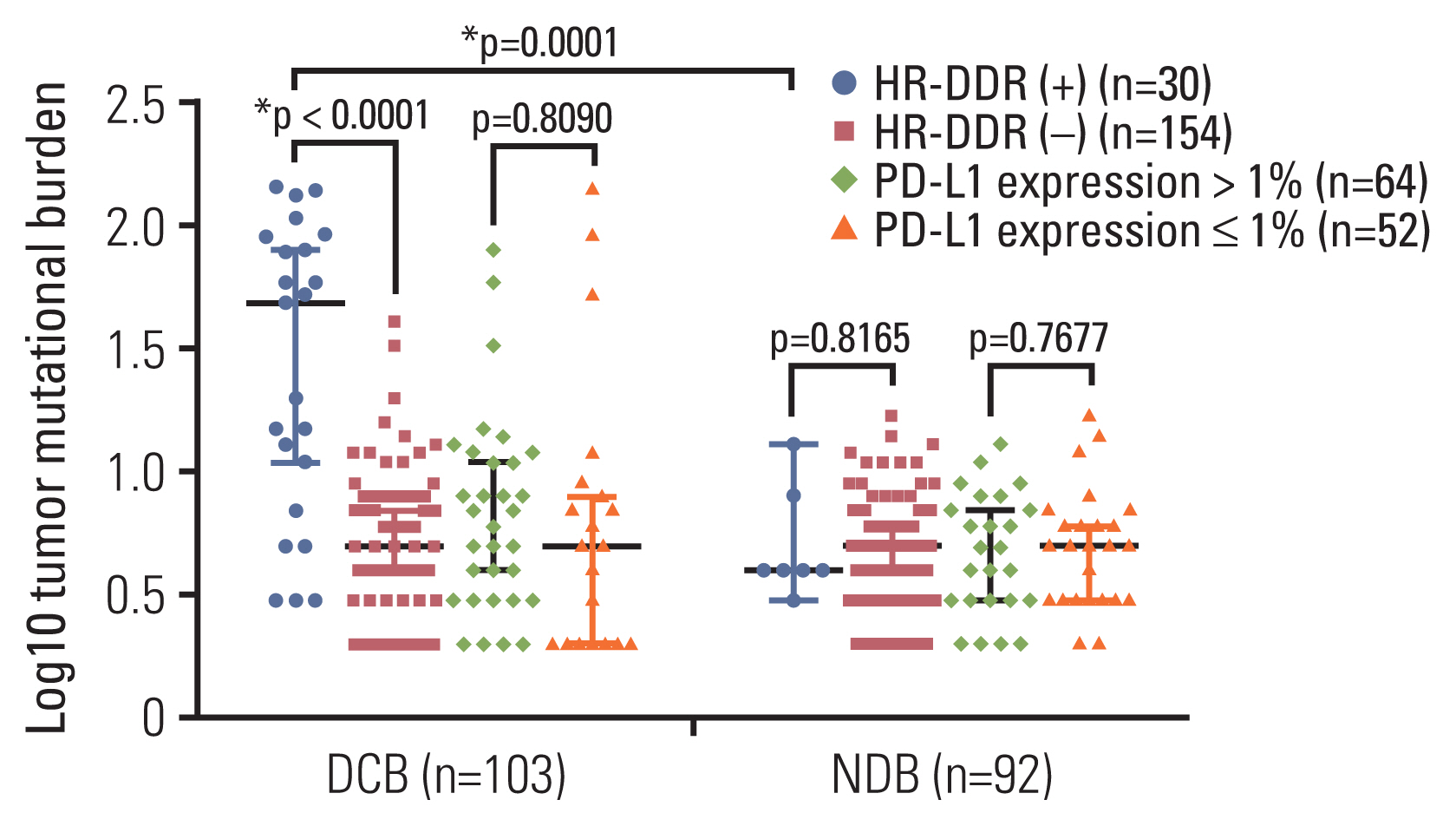
Table 1Baseline characteristics of 1,740 and 592 patients across multiple tumor types sequenced with 295- and 1021-gene panels, respectively
Table 2The characteristics of patients administrated by ICIs treatment
CR, complete response; DCB, durable clinical benefit; HR-DDR, homologous recombination DNA damage repair; ICIs, immune checkpoint inhibitors; NDB, no durable benefit; PD, progression disease; PD-L1, programmed death-ligand 1; PR, partial response; SD, stable disease; TNM, tumor-node metastasis. References1. Jiang T, Shi T, Zhang H, Hu J, Song Y, Wei J, et al. Tumor neoantigens: from basic research to clinical applications. J Hematol Oncol. 2019;12:93.
2. Chen L, Ashe S, Brady WA, Hellstrom I, Hellstrom KE, Ledbetter JA, et al. Costimulation of antitumor immunity by the B7 counterreceptor for the T lymphocyte molecules CD28 and CTLA-4. Cell. 1992;71:1093–102.
3. Leach DR, Krummel MF, Allison JP. Enhancement of antitumor immunity by CTLA-4 blockade. Science. 1996;271:1734–6.
4. Larsen TV, Hussmann D, Nielsen AL. PD-L1 and PD-L2 expression correlated genes in non-small-cell lung cancer. Cancer Commun (Lond). 2019;39:30.
5. Hirano F, Kaneko K, Tamura H, Dong H, Wang S, Ichikawa M, et al. Blockade of B7-H1 and PD-1 by monoclonal antibodies potentiates cancer therapeutic immunity. Cancer Res. 2005;65:1089–96.
6. Topalian SL, Hodi FS, Brahmer JR, Gettinger SN, Smith DC, McDermott DF, et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N Engl J Med. 2012;366:2443–54.
7. Motzer RJ, Escudier B, McDermott DF, George S, Hammers HJ, Srinivas S, et al. Nivolumab versus everolimus in advanced renal-cell carcinoma. N Engl J Med. 2015;373:1803–13.
8. Bracarda S, Altavilla A, Hamzaj A, Sisani M, Marrocolo F, Del Buono S, et al. Immunologic checkpoints blockade in renal cell, prostate, and urothelial malignancies. Semin Oncol. 2015;42:495–505.
9. Brahmer JR, Tykodi SS, Chow LQ, Hwu WJ, Topalian SL, Hwu P, et al. Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. N Engl J Med. 2012;366:2455–65.
10. Reck M, Rodriguez-Abreu D, Robinson AG, Hui R, Csoszi T, Fulop A, et al. Pembrolizumab versus chemotherapy for PD-L1-positive non-small-cell lung cancer. N Engl J Med. 2016;375:1823–33.
11. Le DT, Durham JN, Smith KN, Wang H, Bartlett BR, Aulakh LK, et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science. 2017;357:409–13.
12. Folprecht G. Tumor mutational burden as a new biomarker for PD-1 antibody treatment in gastric cancer. Cancer Commun (Lond). 2019;39:74.
13. Shekarian T, Valsesia-Wittmann S, Brody J, Michallet MC, Depil S, Caux C, et al. Pattern recognition receptors: immune targets to enhance cancer immunotherapy. Ann Oncol. 2017;28:1756–66.
14. Li A, Yang JJ, Zhang XC, Zhang Z, Su J, Gou LY, et al. Acquired MET Y1248H and D1246N mutations mediate resistance to MET inhibitors in non-small cell lung cancer. Clin Cancer Res. 2017;23:4929–37.
15. Buchhalter I, Rempel E, Endris V, Allgauer M, Neumann O, Volckmar AL, et al. Size matters: Dissecting key parameters for panel-based tumor mutational burden analysis. Int J Cancer. 2019;144:848–58.
16. Heeke AL, Pishvaian MJ, Lynce F, Xiu J, Brody JR, Chen WJ, et al. Prevalence of homologous recombination-related gene mutations across multiple cancer types. JCO Precis Oncol. 2018;2018:PO.1700286.
17. Rimm DL, Han G, Taube JM, Yi ES, Bridge JA, Flieder DB, et al. A prospective, multi-institutional, pathologist-based assessment of 4 immunohistochemistry assays for PD-L1 expression in non-small cell lung cancer. JAMA Oncol. 2017;3:1051–8.
18. Campesato LF, Barroso-Sousa R, Jimenez L, Correa BR, Sabbaga J, Hoff PM, et al. Comprehensive cancer-gene panels can be used to estimate mutational load and predict clinical benefit to PD-1 blockade in clinical practice. Oncotarget. 2015;6:34221–7.
19. Ma F, Guan Y, Yi Z, Chang L, Li Q, Chen S, et al. Assessing tumor heterogeneity using ctDNA to predict and monitor therapeutic response in metastatic breast cancer. Int J Cancer. 2020;146:1359–68.
20. Wang Y, Zhao C, Chang L, Jia R, Liu R, Zhang Y, et al. Circulating tumor DNA analyses predict progressive disease and indicate trastuzumab-resistant mechanism in advanced gastric cancer. EBioMedicine. 2019;43:261–9.
21. Zhou Z, Zhu L, Niu X, Shen S, Zhao Y, Zhang J, et al. Comparison of genomic landscapes of large cell neuroendocrine carcinoma, small cell lung carcinoma, and large cell carcinoma. Thorac Cancer. 2019;10:839–47.
22. Zhang Y, Chang L, Yang Y, Fang W, Guan Y, Wu A, et al. The correlations of tumor mutational burden among single-region tissue, multi-region tissues and blood in non-small cell lung cancer. J Immunother Cancer. 2019;7:98.
23. Sun S, Liu Y, Eisfeld AK, Zhen F, Jin S, Gao W, et al. Identification of germline mismatch repair gene mutations in lung cancer patients with paired tumor-normal next generation sequencing: a retrospective study. Front Oncol. 2019;9:550.
24. Singal G, Miller PG, Agarwala V, Li G, Kaushik G, Backenroth D, et al. Association of patient characteristics and tumor genomics with clinical outcomes among patients with non-small cell lung cancer using a clinicogenomic database. JAMA. 2019;321:1391–9.
25. Yao Y, Zhang T, Qi L, Liu R, Liu G, Wang X, et al. Competitive endogenous RNA network construction and comparison of lung squamous cell carcinoma in smokers and nonsmokers. Dis Markers. 2019;2019:5292787.
26. Wang K, McDermott JD, Schrock AB, Elvin JA, Gay L, Karam SD, et al. Comprehensive genomic profiling of salivary mucoepidermoid carcinomas reveals frequent BAP1, PIK3CA, and other actionable genomic alterations. Ann Oncol. 2017;28:748–53.
27. Chalmers ZR, Connelly CF, Fabrizio D, Gay L, Ali SM, Ennis R, et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med. 2017;9:34.
28. Salem ME, Puccini A, Grothey A, Xiu J, Goldberg R, Kim ES, et al. Comparative molecular analysis between microsatellite instability-high (MSI-H) tumors with high tumor mutational burden (TMB-H) versus MSI-H tumors with TMB-intermediate/low. Ann Oncol. 2018;29(Suppl 8):VIII649–69.
29. Pursell ZF, Isoz I, Lundstrom EB, Johansson E, Kunkel TA. Yeast DNA polymerase epsilon participates in leading-strand DNA replication. Science. 2007;317:127–30.
30. Petitjean A, Mathe E, Kato S, Ishioka C, Tavtigian SV, Hainaut P, et al. Impact of mutant p53 functional properties on TP53 mutation patterns and tumor phenotype: lessons from recent developments in the IARC TP53 database. Hum Mutat. 2007;28:622–9.
31. Yang S, Yu X, Fan Y, Shi X, Jin Y. Clinicopathologic characteristics and survival outcome in patients with advanced lung adenocarcinoma and KRAS mutation. J Cancer. 2018;9:2930–7.
32. Telli ML, Timms KM, Reid J, Hennessy B, Mills GB, Jensen KC, et al. Homologous Recombination Deficiency (HRD) score predicts response to platinum-containing neoadjuvant chemotherapy in patients with triple-negative breast cancer. Clin Cancer Res. 2016;22:3764–73.
33. Teo MY, Seier K, Ostrovnaya I, Regazzi AM, Kania BE, Moran MM, et al. Alterations in DNA damage response and repair genes as potential marker of clinical benefit from PD-1/PD-L1 blockade in advanced urothelial cancers. J Clin Oncol. 2018;36:1685–94.
34. Kim SJ, Kim S, Kim DW, Kim M, Keam B, Kim TM, et al. Alterations in PD-L1 expression associated with acquisition of resistance to ALK inhibitors in ALK-rearranged lung cancer. Cancer Res Treat. 2019;51:1231–40.
35. Rosenberg JE, Hoffman-Censits J, Powles T, van der Heijden MS, Balar AV, Necchi A, et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet. 2016;387:1909–20.
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||