The Generation and Application of Patient-Derived Xenograft Model for Cancer Research
Article information
Abstract
Establishing an appropriate preclinical model is crucial for translational cancer research. The most common way that has been adopted by far is grafting cancer cell lines, derived from patients. Although this xenograft model is easy to generate, but has several limitations because this cancer model could not represent the unique features of each cancer patient sufficiently. Moreover, accumulating evidences demonstrate cancer is a highly heterogeneous disease so that a tumor is comprised of cancer cells with diverse characteristics. In attempt to avoid these discrepancies between xenograft model and patients’ tumor, a patient-derived xenograft (PDX) model has been actively generated and applied. The PDX model can be developed by the implantation of cancerous tissue from a patient’s tumor into an immune-deficient mouse directly, thereby it preserves both cell-cell interactions and tumor microenvironment. In addition, the PDX model has shown advantages as a preclinical model in drug screening, biomarker development and co-clinical trial. In this review, we will summarize the methodology and applications of PDX in detail, and cover critical issues for the development of this model for preclinical research.
Introduction
According to the report of World Health Organization and National Cancer Institute, cancer is defined as a group of diseases involving abnormal cell growth with the potential to invade or spread to other parts of the body. In 2012, an estimated 14.1 million new cancer cases and 8.2 million cancer deaths occurred worldwide [1], that comprises 14.6% of total human deaths. Cancer is a leading cause of death worldwide in countries of all income levels. The number of cancer cases and deaths is expected to grow rapidly as populations grow, age, and adopt lifestyle behaviors that increase cancer risk [2]. In this aspect, cancer research needs to be performed in depth based on the model that better represents the exact characteristic of human cancer.
Cancer cell lines (CCLs) derived from patients have been used for not only in vitro but also in vivo experiments [3]. Especially, a xenograft model generated by the injection of these cell lines subcutaneously into immunodeficient mice is the most commonly used model in preclinical drug development [4]. This model is easy to generate and shows consistent tumor growth among animals. However, the xenograft model using CCLs has several limitations on the other hand. The first one is this model could not reflect the patient’s drug response sufficiently. The fact that successful clinical approval rate for cancer drugs is very low (approximately lower than 15%), especially in solid tumor [5] supports this idea. Second issue comes from an observation that most tumors contain highly heterogeneous subsets of cancer cells with different characteristics in one tumor. This is partly due to the genomic instability of cancer that increases the complexity of the cancer phenotype by the simultaneous alterations in several oncogenes and tumor suppressors [6]. Therefore, the xenograft model using CCL is not sufficient to represent the complex tumor heterogeneity. Moreover, tumor cell is often surrounded by fibroblast, vessel cells, and immune cells and interaction with these cells is a critical factor for the characteristics of each tumor. For example, this tumor microenvironment affects the growth and metastasis of cancer cells. However, the xenograft tumor often cannot recapitulate the tumor microenvironment.
To overcome these limitations, there has been increasing interest in patient-derived xenograft (PDX) model as a more advanced preclinical cancer model. In the PDX model, a tumor specimen is directly transplanted into immunodeficient mice, providing a faithful representation of individual tumor [7]. Although genetically engineered mouse model is also generally used as a preclinical model, it usually takes long time to develop and has less heterogeneity than PDX model because only few genes are modified. Therefore, PDX model is becoming a preferred preclinical tool in the drug development [8,9]. The translational cancer research is also facilitated by several collections of extensively characterized PDX model and it is regarded as a useful tool to apply the personalized medicine. In this review, we will cover current methodology for the generation of PDX models, summarize PDX model developed so far, describe the advantage of PDX model, and comment important issues for the future development of this model.
Methodology for Generating and Validating the PDX Model
1. The process of generating PDX
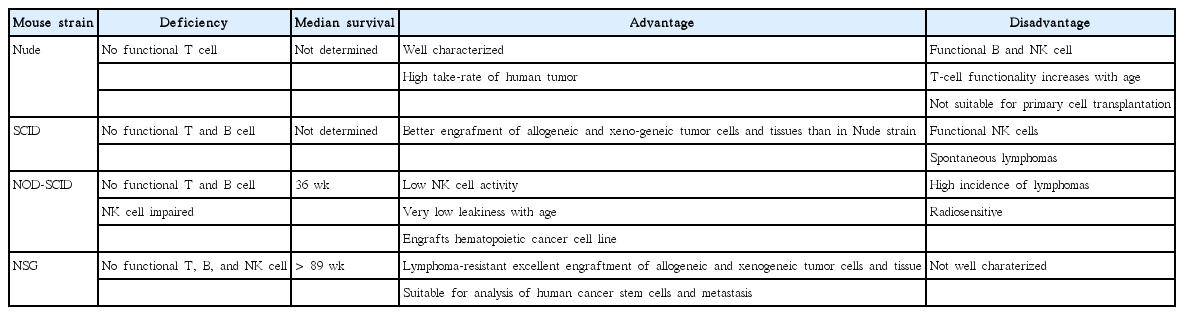
PDX can be developed by the implantation of cancerous tissue from a patient’s tumor into an immunodeficient mouse directly. Several types of such mice can be used to establish PDX models: athymic nude mice, severely compromised immune deficient (SCID) mice, nonobese diabetic (NOD)– SCID mice, and recombination-activating gene 2 (Rag2)- knockout mice [10]. The mice used must be “immunocompromised,” to prevent graft rejection. The NOD-SCID mouse is more commonly used for PDX models than SCID because it does not produce natural killer cells [9]. However, recently, NSG (NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ), mice are more preferred than any other strains. NSG has a complete null mutation in the gene encoding the interleukin 2 receptor gamma chain, which causes deficiency in multiple cytokine signaling pathway, leading to dysfunctions of many parts of innate immunity including natural killer cell differentiation [11]. Therefore, it is regarded as best model for the efficient engraftment of primary human tumor or tumor cells [12,13]. The advantages and disadvantages of each strain are summarized in Table 1.
There are two different input materials for the PDX generation. The first one is single-cell suspension and the second one is tissue fragment. Usually implanted tumor is gained when patient underwent surgery. The lesion of cancer is transported to pathologist or researcher in tube filled with culture media (Dulbecco's modified Eagle's medium [DMEM] or RPMI with 10% fetal bovine serum [FBS] and 1% penicillin/streptomycin). After that, the necrotic tissues are removed and the tumors are sectioned into smaller fragments, chemically digested, or physically manipulated into a single-cell suspension. There are advantages and disadvantages in utilizing either discrete tumor fragments or singlecell suspensions. Tumor fragments retain cell-cell interactions as well as some tissue architecture of the original tumor, therefore mimicking the tumor microenvironment. Alternatively, a single-cell suspension enables scientists to collect an unbiased sampling of the whole tumor, avoiding spatially enriched subclones that are otherwise inadvertently selected during analysis or tumor passaging [14]. However, singlecell suspensions subject tumor cells to harsh chemical or mechanical forces that may sensitize cells to anoikis, taking a toll on cell viability and engraftment success [15].
Unlike developing xenograft models using CCLs, there are no intermediate in vitro processing steps before implanting tumor fragments into mouse to create a PDX. When using CCLs, a sufficient amount of cells should be cultured before the injection into mouse. This in vitro step consumes large amount of time and effort. However, in the case of PDX, tumor fragments can be implanted immediately after obtaining tissue from patient without any culture. Instead, tumor fragments should be chopped into 2-3 mm3 for easier transplantation (Fig. 1). The tumor fragments are either implanted heterotopically or orthotopically onto the recipient mouse. Heterotopical implants occur when the tumor fragment is implanted into an area of the mouse unrelated to the original tumor site, generally subcutaneously or subrenal capsular sites [16]. In the orthotopic transplant, in contrast, scientists transplant the patient’s tumor tissue into the corresponding anatomical organ. Subcutaneous PDX rarely produce metastasis in mice and hardly simulate the initial tumor microenvironment [16]. Orthotopic model is difficult to generate, depending on organ, but it can mimic the natural environment of primary tumor, so it is thought as an ideal model. Ultimately, it takes about 2 to 4 months for the tumor to engraft. The latency varied by tumor type, implant location, and strain of immunodeficient mice utilized. The engraftment failure should not be declared until no tumor growth is observed for, at least, 6 months [10].
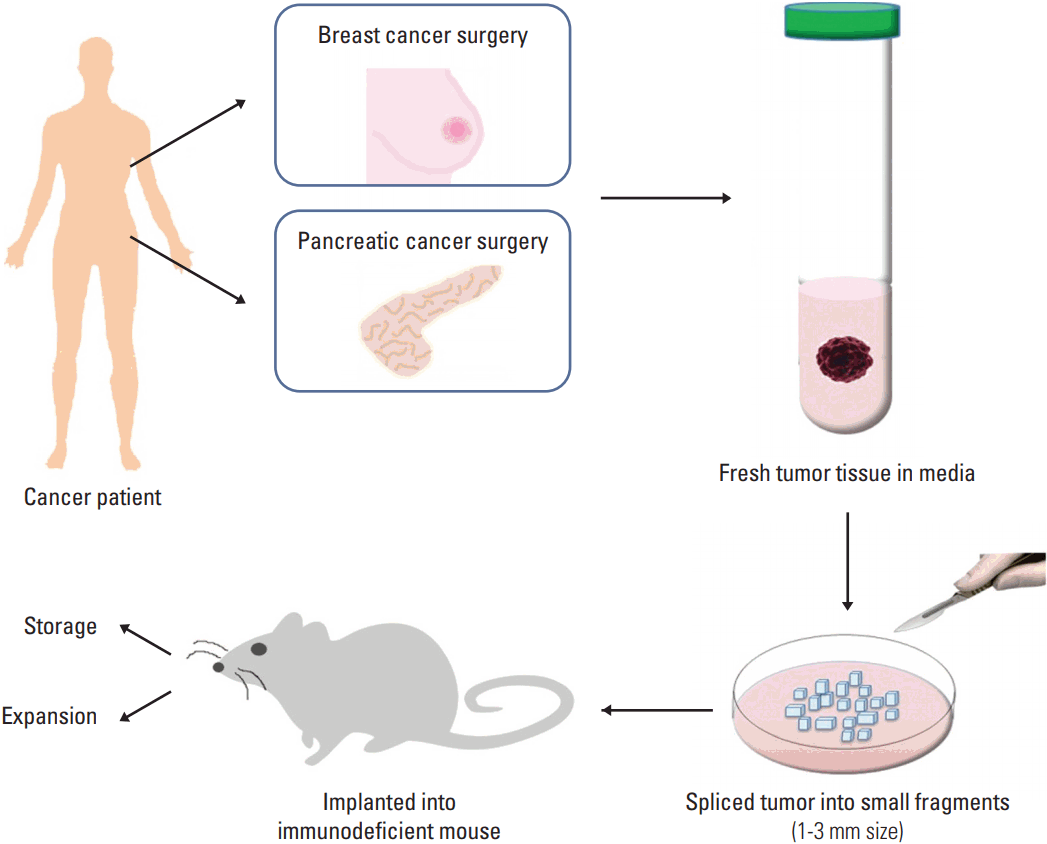
An overall procedure for the generation of patient-derived xenograft (PDX) model. The PDX model can be developed by the implantation of fragments from a patient’s tumor into an immunodeficient mouse directly. A part of tumor from surgery (breast or pancreatic tumor in this example) is put in preserving media and the tumor is sliced into small fragments. The fragments are implanted subcutaneously or into the orthotopical organ, for example, mammary fat pad in the case of breast cancer.
After tumor is harvested, most parts of tumor are cut into small pieces (1-2 mm3) and washed 3-4 times with culture media. Finally, the tissue is put in freezing media (FBS plus 10% dimethyl sulfoxide) and stored at -80℃. This stock can be used for re-transplantation into other mice later. The first generation of mice receiving the patient's tumor fragments are commonly denoted F0. When the tumor reached enough size, it is dissected and re-implanted into other recipient mice. Each generation thereafter is denoted F1, F2, F3, ..., and Fn. For drug development studies, PDX underwent more than F3 generation is often used, after ensure that the PDX has not genetically or histologically diverged from the original tumor [17]. When tumor saved at -80℃ is re-transplanted, the stock is thawed completely in 37℃ water bath and the tissue is put in plastic tube filled with 10-15 mL media (DMEM or RPMI containing 1% penicillin/streptomycin). These fragments of tumor are moved to petri dish for re-implantation.
2. Success rate for the generation of PDX model
Some companies have commercialized PDX model for almost all cancer types. Depending on the cancer type, the success rate of the xenograft varies. Based on previous reports, success rate and specific engraftment method are summarized in Table 2 [18-33]. The success rates of PDX are influenced by several factors. Firstly, the characteristics of primary tumor, such as the aggressiveness of tumor, histological type, and tumor cell percentage in the tissue, are considered important. For example, in the case of breast cancer, triple negative breast cancer shows higher success rate than other histological type like estrogen receptor-positive or human epidermal growth factor receptor 2-positive cancer [34]. Secondly, process for the generation of PDX can affect the success rate. Tissue should be as fresh as possible, meaning that time taking from surgery room to laboratory should be minimized. Also, the tissue needs to be kept in cold, fresh media immediately after surgery. Thirdly, the size and number of tissue implanted can be a factor affecting success rate. The greater number of pieces tends to increase success rate and fragment size should be 1-2 mm3. Fourthly, the location of implantation can make difference in success rate. Usually, implantation in subrenal capsule shows higher success rate than subcutaneous implantation because subrenal capsule maintains the original tumor stroma as well as the equivalent host stroma [35]. Notably, the ability of tumor to adapt to host environment varies by the original tumor. Tissues that adapt well to environment without immune rejection are more likely to succeed.
3. Validation and molecular analysis of PDX model
When the transplanted tumor reaches enough size (diameter of 1-2 cm), it should be harvested. During the harvest process, mouse tissue surrounding tumor should be removed as much as possible. After that, a portion of tumor is preserved in 10% formalin. To validate that the gross histology of the PDX is conserved with the original tumor, a paraffin block will be generated from the preserved tissue and will be stained by hematoxylin and eosin or other important immunohistochemical markers.
On the other hand, genomic DNA extracted from frozen tissue would be used for cancer panel or whole genome sequencing. RNA also would be purified for RNA sequencing or real-time polymerase chain reaction to measure the expression of specific genes. If the expression of proteins needs to be examined, protein can be extracted from the PDX tissue by chopping or grinding it with tissue-rupter.
Application of PDX for Cancer Research
1. Advantages of using PDX model
CCLs are originally derived from patient tumors, but has been extensively selected and changed to acquire the ability to proliferate in vitro. Because of the in vitro manipulation, CCL also gained genetic transformations that are not recovered when cells grow in vivo [36]. As a result of the process including enzymatic dissociation and centrifugation, cells that are adapted to survive in culture are phenotypically homogeneous [14]. Consequently, the tumor generated by injection of CCL is unlikely to retain the heterogeneity of its original tumor.
Researchers deliberated the reason why only 5% of anti-cancer agents are approved by the Food and Drug Administration after pre-clinical testing. As a result, the lack of tumor heterogeneity and the absence of the human stromal microenvironment affect high rate of failure [37]. Specifically, CCL-xenografts often are not predictive for the drug response of the primary tumors because it lacks the heterogenous molecular pathways of drug resistance existing in original tumor or the other cells comprising tumor microenvironment [37]. In other words, drugs usually target specific molecule or receptor in CCL and work efficiently as it consists of selected homogenous cells. However, in most primary tumors with heterogenous cancer cells with stroma, the response to the drug can be totally different.
To overcome this limitation, PDX models have been successfully established for breast, prostate, pancreas, colorectal, lung, and many other cancers for drug safety and efficacy studies as well as examining personalized response to certain anti-cancer agents [38]. Since PDX can be passaged without in vitro culture steps, PDX models allow the propagation and expansion of patient tumors without significant genetic transformation of tumor cells over multiple generations [39]. Within PDX models, tumor cells grow in physiologically-relevant tumor microenvironments that mimic the oxygen, nutrient, and hormone levels that are found in the patient’s primary tumor site [17]. Furthermore, implanted tumor tissue maintains the genetic and epigenetic abnormalities found in the patient and the xenograft tissue can be excised from the patient to include the surrounding human stroma [40]. As a result, numerous studies have found that PDX models exhibit similar responses to anti-cancer agents as seen in the actual patient who provided the tumor sample [41]. Hence, PDX model is beneficial to test therapeutic responses for certain drugs because multiple therapies can be tested against expanded PDXs generated from one biopsy and pretreatment and post-treatment data can be acquired from the xenograft tissues [40].
2. Application of PDX model in cancer research
1) Drug screening and biomarker development
PDX model has been generated and used in several retrospective studies and more recently in preclinical trials. Several researches using PDX model in breast cancer, renal cell cancer, non-small cell lung cancer (NSCLC), squamous cell carcinoma of the head and neck, and colorectal cancer showed that the drug response rates in PDX models correlate with those observed in the clinic, both for targeted agents and for classic cytotoxins. For example, in renal cell cancer, PDX model respond to the sirolimus and sunitinib and dovitinib, but not to erlotinib, which is the same as clinical data [42]. In addition, the response rate of epidermal growth factor receptor inhibitor cetuximab in PDX model was exactly identical with the results from clinic [18]. Similarly, an efficacy study for MEK and phosphoinositide 3-kinase/mammalian target of rapamycin inhibitor in PDX model showed low response, which also coincide with clinical study [43]. With regard to conventional chemotherapy, studies in NSCLC, breast cancer, colorectal cancer, and pancreatic ductal adenocarcinoma demonstrated that PDX model showed comparable response rates for clinically used agents such as paclitaxel, carboplatin, gemcitabine, 5-fluorouracil, irinotecan, and adriamycin to clinical data [44].
More recently, the use of PDX models as potential screening platforms for clinical trials has been shown in a prospective study for pancreatic cancer. In PDX model, metformin treatment did not inhibit the growth of pancreatic cancer. Clinical trial also showed no benefit of adding metformin to combination therapy regimens targeting locally advanced and metastatic pancreatic cancer [45]. This is an example showing the data from preclinical test using PDX model could save time and resource required for clinical trials.
With regards to the discovery of biomarker, the concordance between PDX models and human trials allows us to discover biomarker for drug susceptibility and drug resistance. Furthermore, a relationship between drug efficacy and molecular (genetic) characteristics can be easily studied by using heterogeneous PDX models. For example, Sebastiani et al. [46] revealed relationship to molecular mechanism of gemcitabine resistance and survival in pancreatic cancer. PDX models are also valuable tools for generating drugresistant tumor models, which is achieved by repeated administration of a drug. Das Thakur et al. [47] generated vemurafenib-resistant melanoma model using PDX. They found elevated mutant BRAF protein by continuous drug treatment and this mutation is a critical factor for vemurafenib resistance. Likewise, proteomic or genetic comparison between sensitive and resistant models can be explored for the identification of prognostic biomarkers in clinical studies.
2) Co-clinical trial
The co-clinical trial denotes a clinical trial conducted in parallel with preclinical trial. The relevant clinical, biological, and pharmacologic information (i.e., somatic mutational background, germline single nucleotide polymorphism variations, responsiveness to specific regimens, imaging, microarray data, and proteomics profiles) are analyzed comprehensively, and integrated to identify biomarkers that predict a response to specific treatment [48-50]. In this approach, PDX model is developed from a patient enrolled in clinical trial and the PDX is treated with the same regimen to monitor clinical response [44]. By establishing PDXs of a patient enrolled in a clinical trial and treating it with a new agent, a screening for prognostic biomarkers can be performed and underlying drug response mechanisms can be studied. Based on this, novel combination strategies can be proposed. For example, Heid et al. [51] performed co-clinical assessment of tumor cellularity in pancreatic cancer. Owonikoko et al. [52] also reported that PDX faithfully replicated clinical outcome in a phase II co-clinical trial of arsenic trioxide in relapsed small cell lung cancer. Although it is not commonly tried so far, the co-clinical trial may be increased in the future because it can save time for drug development and reflects personalized medicine in a preclinical setting.
3) Precision medicine
Oncology research has evolved in parallel with the improved understanding of the cancer genotype and phenotype, leading to a new era of precision medicine [53]. This is colloquially termed “the right drug, for the right patient, at the right time.” In contrast to the conventional chemotherapy, the precision medicine combines individual patient’s characteristics, i.e., the genomic landscape of each tumor with molecularly targeted agents or immunotherapeutics to maximize treatment efficacy and minimize side effects [53]. Practically, the concept of precision medicine is to group patients into subpopulation based on sophisticated genomic profiling, enabling certain therapies to target the subgroup specifically [6]. Based on this notion, PDX is an appropriate model as it retains the genomic characters of individual tumor and represents a subgroup with similar genetic profile. Moreover, PDX model can even recapitulate heterogeneity within the same tumor specimen (intratumoral heterogeneity).
For example, a recent report showed integrated PDX models delineate individualized therapeutic vulnerabilities of pancreatic cancer [54]. In this study, an exom sequencing of patient tumor revealed multiple conserved genetic alterations. Using the PDX model, they showed the effectiveness and selectivity of treatment responses to more than 500 single and combination drug regimens. In addition, a genomic information-driven treatment was performed in advanced breast cancer using targeted next-generation sequencing (NGS), array-based comparative genomic hybridization data obtained from PDX model [55]. In ovarian cancer, anti-cancer drugs were tested for precision medicine approach, using PDX model [56]. Also, fine-tuning PDX model for precision medicine was tried in consideration of cross-species cytokine in leukemia [57]. Collectively, the efforts to implement personalized medicine using PDX model (summarized in Fig. 2) has been made in various cancers and enormous data would be accumulated with the progress of NGS and bioinformatic techniques [3].

A flowchart showing the establishment of personalized medicine using patient-derived xenograft (PDX) model. Genomic signature of a primary tumor is analyzed by next-generation sequencing. At the same time, tumor fragments are implanted into immunodeficient mouse. The patient will be treated with the drug that showed best response in PDX. Also, a database of integrated genomic signature would be established to predict a drug response for a new patient with similar genomic signature. SNP, single nucleotide polymorphism.
Limitations and Challenges in PDX Model
Although PDX model is relatively easy and ideal model to research the cancer, there are limitations and challenges that need to be overcome. Firstly, the most appropriate tissue should be transplanted into mouse. In the case when a tumor size is very large, the part that can represent the tumor should be used for PDX. In this step, the help of pathologist and surgeon would be needed. Moreover, the tissue should be as fresh as possible, to generate PDX efficiently. Sometimes, smaller sample, such as fine-needle aspirations, are required to be transplanted for personalized medicine application. Therefore, the PDX technique using small specimen should be more studied and developed. Secondly, it is important to define the best strategy of engraftment in mice for different tumor type. Orthotopic implantation is considered more ideal, but special surgical technique is required depending on the cancer type and this way takes much time and effort for development. Subcutaneous implantation has relatively higher success rate and simpler procedure, so this approach is useful when lots of mouse needed to be grafted in the short time. Thirdly, duration of survival and treatment schedules for patients is also a considerable factor for personalized medicine applications. It normally takes 2-8 months to develop a PDX model for a preclinical study. In many cases, however, this length of time is too long for patients to wait. Fourthly, we need to deal with engraftment failure that is still high for some tumor types, such as hormone receptor–positive human breast cancer. Fifthly, a key aspect in PDX research is the use of immunodeficient mouse strain for tumor engraftment and propagation because mouse should avoid rejection of human tissue. For this reason, conventional PDX model is not appropriate for the screening of immune-modulating agents, such as vaccines and checkpoint blocking antibodies that function by activating host immune system. Instead, a humanized PDX model that has been transplanted human hematopoietic stem cells (CD34+ cells) [58] can be an alternative option. Sixthly, the replacement of human stromal components (such as cancerassociated fibroblasts, endothelial cells, immune and inflammatory cells) by murine elements could happen with repeated passages [59]. In this regard, the PDX has limit to mimic totally exact tumor microenvironment of human cancer and needs to find the way to overcome it.
Lastly, financial aspect cannot be ignored. It needs a substantial cost to develop PDX model because the immunodeficient mice are expensive and it takes long time to develop tumor. Nevertheless, the PDX is still considered as a worthy preclinical tool because of its uniqueness and many advantages as mentioned above.
Conclusion
As described in this review, the PDX model retains tumor characteristics such as cell-cell interactions and tumor microenvironment, although with some limitations. It is becoming clear that this model has many advantages in preclinical tests of drug screening, biomarker development and co-clinical trial. With the advances of NGS and other omics technique, we are approaching to obtain a comprehensive molecular signature for each patient’s tumor. We believe the PDX model is one of the appropriate preclinical tools to widen the personalized medicine strategy in the future. More studies for improving graft success rate as well as generating humanized PDX with the patients’ immune cells will accelerate the usage of this preclinical model.
Notes
Conflict of interest relevant to this article was not reported.
Acknowledgements
This work is supported by a grant of the Basic Science Research Program through a grant from Asan Institute for Life Sciences, Asan Medical Center (No. 2017-571) and a grant of Korean Health Technology R&D Project, Ministry of Health & Welfare, Republic of Korea (Grant Number: H14C2640).