Nucleotide Excision Repair Gene ERCC2 and ERCC5 Variants Increase Risk of Uterine Cervical Cancer
Article information
Abstract
Purpose
Defects in the DNA damage repair process can cause genomic instability and play an important role in cervical carcinogenesis. The purpose of this study was to analyze the association of 29 candidate single nucleotide polymorphisms (SNPs) in genes in the DNA repair pathway, TP53, and TP53BP1 with the risk of cervical cancer.
Materials and Methods
Twenty-nine SNPs in four genes in the DNA repair pathway (ERCC2, ERCC5, NBS1, and XRCC1), TP53, and TP53BP1 were genotyped for 478 cervical cancer patients and 922 healthy control subjects, and their effects on cervical carcinogenesis were analyzed.
Results
The most significant association was found for rs17655 in ERCC5, with an age-adjusted p-value < 0.0001, for which a strong additive effect of the risk allele C was observed (odds ratio, 2.01 for CC to GG). On the other hand, another significant polymorphism rs454421 in ERCC2 showed a dominant effect (odds ratio, 1.68 for GA+AA to GG) with an age-adjusted p-value of 0.0009. The association of these polymorphisms remained significant regardless of the age of onset. The significant result for rs17655 was also consistent for subgroups of patients defined by histology and human papillomavirus (HPV) types. However, for rs454421, the association was observed only in patients with squamous cell carcinoma and non-HPV 18 type.
Conclusion
The results of this study show a novel association of cervical cancer and the genes involved in the nucleotide excision pathway in the Korean population.
Introduction
The incidence of invasive cervical cancer has been decreasing (annual percent change [APC] in age-adjusted rates of –4.2%), however cervical neoplasm is still a major health problem in Korea. According to the National Cancer Registration statistics in Korea, the age-standardized incidence rate of cervical cancer was 14.9 per 100,000 in 2011, and the age standardized incidence rate for cervical carcinoma in situ increased from 7.5 in 1993 to 19.0 per 100,000 females in 2009 (APC, 5.7; 95% confidence interval [CI], 4.8 to 6.7) [1]. The incidence of adenocarcinoma has remained stable despite the decreased incidence of squamous cell carcinoma. Eighty percent of infections of human papillomavirus (HPV) clear within 3 years, and persistence of viral DNA is reported in 5%-15% of infected women [2]. Finally, only 3%-5% of women with persistent HPV infection reach the stage of cervical intraepithelial neoplasm (CIN). With this background, we need to define the subset of the population that is vulnerable to persistent HPV infection and progression to epithelial abnormality. Use of tobacco and estrogen-derivatives are known as external factors associated with progression to cervical cancer [3]. However, reports of familial aggregation of cervical cancer from Swedish and Latin American cohort studies suggest that host genetic factors may contribute to cervical carcinogenesis, although shared environmental factors in childhood may be partly responsible for familial aggregation [4,5].
In cervical cancer, HPV integration into the genome of host cells initiates E6 and E7 oncoprotein expression, which in turn drives differentiating cells into S or G2/M phases to induce amplification of the viral genome. Once viral genetic material is introduced in the host genome, it is considered as "damage," which causes activation of cellular DNA repair machinery [6]. As a result of attempts at unordered replication of cells, formation of multiple sites of stalled replication forks occurs along with activation of DNA damage repair pathways. Activated DNA repair pathways that prevent DNA damage and mutagenesis included in these repair pathways are the base excision repair (BER) pathway, the nucleotide excision repair (NER) pathway, DNA single strand break (SSB), and the double strand break repair (DSB) pathways [7]. X-ray repair cross-complementing (XRCC) 1 in the BER pathway and excision repair cross-complementing (ERCC) 2 and ERCC5 in the NER systems were previously studied as risk factors for epithelial cancer [8]. Viral E6 and E7 activate the ataxia telangiectasia mutated (ATM) DNA damage response necessary for the differentiation-dependent amplification of viral genomes [9]. TP53 is a major activator of ATM upon DNA damage signal, and functional polymorphisms in TP53 and TP53BP1 are known to be associated with genetic instability as well as cancer risk [10].
Several common and putative functional single nucleotide polymorphisms (SNPs) of these DNA repair genes were reported as genetic factors associated with HPV persistence and cervical cancer progression [11]. In the current study, 29 genetic polymorphisms associated with four genes in the DNA repair pathways, TP53 and TP53BP1 were examined to determine their effects on development of cervical cancer.
Materials and Methods
1. Study population
The study was designed as a hospital based case-control study. A total of 479 cervical cancer patients who visited the National Cancer Center, Korea between July 2003 and March 2011 were recruited. The control group was selected from a cohort recruited at the Center for Early Detection and Prevention at the same hospital between August 2002 and August 2011. A total of 8,291 women who provided informed consent completed a questionnaire to collect information on demographics and cancer risk factors, and provided blood samples; 4,005 women had DNA samples available in an adequate amount (≥ 10 μg). Among them, 1,000 consecutive control subjects with informed consent were selected in chronological order starting from August 2002. Those who had missing genotypes due to inadequate DNA quality were excluded (1 case and 4 controls); those with history of other types of cancer (72 subjects) were also excluded from control samples. As a result, 478 cases and 922 control subjects were used in the analysis. The study was approved by the Institutional Review Board of the National Cancer Center in Korea (IRB No. NCCNCS 10-412).
2. Selection and genotyping of 29 SNPs
After an intensive literature review, six genes (ERCC2, ERCC5, NBS1, TP53, TP53BP1, and XRCC1) involved in the DNA damage repair pathway were selected. The candidate polymorphisms in the six genes, including eight non-synonymous SNPs, were selected on the basis of previously reported associations with cancer risk or prognosis. Five SNPs of TP53 and TP53BP1 were selected because they are frequently studied as cancer predisposing factors. The minor allele frequency (> 5%) in the Asian population was an additional filter for selection of target SNPs. Genomic DNA was extracted from peripheral blood samples using a QIAamp DNA Blood Mini Kit (Qiagen, Valencia, CA) in accordance with the manufacturer’s instructions. The TaqMan-Allelic Discrimination method was used for genotyping SNPs [12]. Primers and probes for the 29 selected SNPs were designed using Primer Express software ver. 2.1 (Applied Biosystems, Foster City, CA). Polymerase chain reaction (PCR) amplifications (5 μL) were performed using 10 ng of template DNA, 1× TaqMan Universal Master Mix (Applied Biosystems), 300 nM of each primer, and 200 nM of each fluorogenic probe in 384-well plates. Thermal cycling was initiated with a 2-minute incubation at 50°C, followed by a first denaturation step of 10 minutes at 95°C, and then 40 cycles of 15 seconds at 95°C and 1 minutes at 60°C. After PCR amplification, the plates were read using an ABI PRISM 7900 Sequence Detection System (Applied Biosystems) and results were analyzed using the Allelic Discrimination software SDS 2.1 program (ver. 5.0, Applied Biosystems).
3. Statistical analysis
The clinical and demographic characteristics of the subjects were compared using a chi-square test for categorical variables and a two-sample t test for continuous variables. The Hardy-Weinberg equilibrium (HWE) for 29 SNPs was examined in the control group.
The association of each marker was analyzed using the minimum p-value of the Pearson’s chi-square test and the Cochran-Armitage trend test based on the additive genetic model (MIN2) [13], and the corresponding p-value associated with MIN2 was obtained [14]. MIN2 proposed by WTCCC has shown robust and efficient power performances over a wide range of underlying genetic models [13,14]. Both crude and age-adjusted analyses were performed for all 29 SNPs by incorporating both age and genotype as exploratory variables in an unconditional logistic regression model. For significant SNPs, subgroup analyses were performed according to the age of onset (≤ 40 or > 40), histology (adenocarcinoma or squamous cell carcinoma), and HPV type (18 vs. non-18). In addition, ordinal outcome variables were formed (case ≤ 40, case > 40, control for age of onset; adenocarcinoma, squamous cell carcinoma, control for histology; HPV 18, non-HPV 18 control for HPV type), and the associations of SNPs with this ordinal outcome were examined using the proportional odds model. A conservative and stringent criterion of the Bonferroni correction was considered in determining the statistical significance at a family-wise error rate of 0.05. As part of the internal validation, a split-sample method was implemented. Using the split-sample approach, data were randomly divided into two equal-sized cohorts; the first cohort served as a training set and the second cohort served as a testing set. In the training set, the association test was performed using MIN2, as previously described, and SNPs with p-values less than 0.1 were further tested using the testing set. SNPs that were significant at the overall type I error rate of 0.05/29 based on the Bonferroni correction were finally selected. Because the results of the split-sample method can be hugely dependent on the random selection of training and testing sets, this procedure was repeated 10 times and the number of significant results obtained out of the 10 repetitions was counted.
Using the bootstrap approach, a thousand data sets were generated using the bootstrap re-sampling procedure. The association was tested, and SNPs with p-values less than 0.05/29 were selected as significant.
Results
1. Patient characteristics
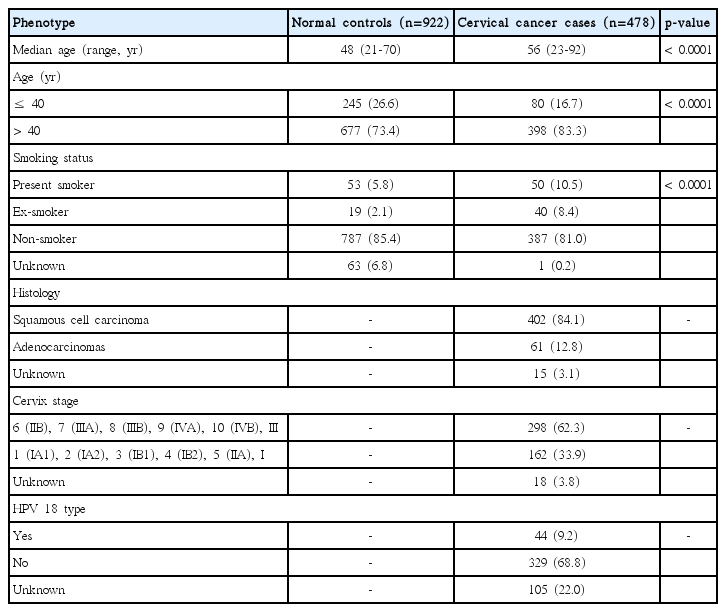
Demographic features of the cases and control subjects are shown in Table 1. Age and smoking status were surveyed in both cases and control subjects. The median age of the case group was higher than that of the control group (median ages of 56 and 48, respectively), and cases were more likely to be current or ex-smokers than control subjects. Among cases, major histologic type was squamous cell carcinoma (84.1%), and advanced stages (≥ IIB) were dominant (62.3%). HPV type 18 was observed in 9.2% of patients.
2. Genetic risk variants for cervical cancer
No deviation from HWE was observed for all 29 SNPs, as evidenced by the high p-value from the HWE test (Supplementary Fig. S1). From the results of the association analysis, four of the six SNPs in the ERCC2 gene, rs454421, rs238417, rs238406, and rs3710366, showed associations with p-value under 0.05 (Supplementary Table 1). Applying a conservative Bonferroni correction (p < 0.05/29), two SNPs of rs454421 and rs17655 were found to show significant association with the risk of cervical cancer. The frequencies of genotypes by case-control status, and the odds ratios under various genetic models for these two SNPs are summarized in Table 2. The most significant association was found for rs17655, which showed a strong additive effect of the risk allele C (age-adjusted p < 0.0001). Subjects carrying the CC genotype showed a 2-fold increased risk for cervical cancer (odds ratio [OR], 2.01) when compared to carriers of the GG genotype. On the other hand, the dominant genotypes of rs454421 (GA+AA) contribute to an increased risk of cervical cancer. In testing for the cumulative effect of these two SNPs, cervical cancer risk was found to increase according to the number of risk alleles, as shown in Table 3. Consistent results were observed when adjusting for both age and smoking status, and when neither was adjusted for. An increased number of risk alleles showed significant association with increased risk of cervical cancer (p for trend < 0.0001). Subjects who carried both risk genotypes showed the highest risk for cervical cancer (OR, 3.75; adjusted p=0.0003).
3. Internal validations using split-sampling and bootstrap
Using the split-sample approach, rs454421 and rs17655 were found to show significant association with the risk of cervical cancer in 9 and 10 split-sample procedures out of 10 repetitions, respectively. In addition, rs454421 and rs17655 were found to show significant association with the disease 781 and 887 times out of 1,000 bootstrap samples, respectively. These results demonstrate the relative robustness of the findings given that no external validation data sets were available.
4. Subgroup analyses of rs454421 and rs17655
The associations of both rs454421 and rs17655 were examined separately in subgroups of patients with early onset (≤ 40) and late onset (> 40) (Supplementary Table 2). Increased ORs of the risk genotypes for both SNPs were consistently observed in both age groups. The ORs of genotypes AA and GA over GG of rs454421 SNP were 1.86 (p=0.0287) and 1.57 (p=0.0011) in the early and late onset groups, respectively. For rs17655, the ORs of GG genotype over CC, and GC over CC, were 1.43 and 2.03 (p=0.0281) in the early onset group, and 1.38 and 1.89 (p=0.0001) in the late onset group. For both SNPs, stronger association in terms of the magnitude of the odds ratio was observed for those in the early onset group, as evidenced by the significant p-values, which are based on the proportional odds model with ordered disease status according to the age of onset (p=0.0002 for rs454421 and p < 0.0001 for rs17655) (Supplementary Table 3). Further subgroup analyses were performed according to histology (squamous cell or adenocarcinomas) and HPV type (18 or non-18) (Supplementary Table 2). This was because our previous study showed significantly lower disease-free survival rates in patients with adenocarcinomas and in patients with HPV 18 genotype [15]. For rs454421, the association was only observed in squamous cell carcinoma (p=0.0004) and non-HPV 18 type (p=0.0043). On the other hand, for rs17655, significant association was found in all subgroups (p-value in squamous cell carcinoma, 0.0002; in adenocarcinoma, 0.0326; p-value in HPV 18 type, 0.0127; in non-HPV 18 type, 0.0009). When multiple testing was adjusted for, similar to the case of rs454421, the association was observed only in squamous cell carcinoma and non-HPV 18 type.
Discussion
DNA repair is an essential process in prevention of genomic alteration caused by carcinogenic stresses. Because DNA repair plays a fundamental role in maintaining genomic integrity, disruption of DNA repair allows genomic instability and cancer development [16]. Several major pathways of DNA repair are involved in the excision of base and nucleotides, in mismatch repair, and in DNA break repair. ERCC groups of ERCC1 (XPA), ERCC2 (XPD), ERCC4 (XPF), and ERCC5 (XPG) are important components in the NER pathway, and have been widely studied in ovarian, colorectal, and non-small cell lung cancers [17-19].
Genetic variants of ERCC genes were also widely studied as risk factors for cancer as well as predictors of treatment outcomes in cancer [8,20]. Our results show highly significant associations of rs454421 in ERCC2 and rs17655 in ERCC5 with cervical cancer. These two SNPs commonly showed a stronger influence in patients who developed cancer after the age of 40, suggesting that other genes may be involved in younger patients with cervical cancer, who have shorter intervals between HPV infection and development of cervical cancer. The effects of the two SNPs on cervical cancer risk were more prominent in squamous carcinoma than in adenocarcinoma. The association of polymorphisms in HPV 18 type was difficult to test due to the small size of the group.
Non-synonymous polymorphism rs17655 (Asp1104His) in ERCC5 has been reported as a risk factor for various cancers, including lung cancer and bladder cancer [8]. It also showed association with increased risk of breast cancer in female radiation technologists who were occupationally exposed to low levels of ionizing radiation [21].
Based on their well-known role as tumor suppressors and DNA damage checkpoints, functional polymorphisms of TP53 and TP53BP1 were also widely studied for various cancer types. Three 53BP1 variant genotypes (Glu353Asp [rs560191], T-885G [rs1869258], and Gln1136Lys [rs2602141]) increased the risk of progesterone receptor–negative breast cancer, and the two latter variants significantly increased estrogen receptor–negative breast cancer in the Chinese population. In addition, TP53BP1 Gln1136Lys (rs2602141) and TP53 Arg72Pro (rs1042522) showed gene-gene interaction with increased breast cancer risk in patients [22].
TP53 codon 72 polymorphism causes different biochemical functions of TP53. The current view is that the TP53-Arg72 protein is more effective at inducing apoptosis and protects cells from tumor development compared with the TP53-Pro72 protein [23]. However, a meta-analysis concluded no association of Pro72Arg variant with incidence of cervical cancer. In addition, Arg variant was previously found to be more susceptible to degradation by the HPV 18 E6 protein [24]. In our subgroup analysis of rs1042522, a tendency toward an association with the risk of cervical cancer was observed for patients with younger age of onset (OR, 1.72; p=0.07), and for patients with HPV 18 infection (OR, 2.09; p=0.03). The small sample size of the subgroup may be the reason that statistical significance was not reached, and further research using larger samples is needed to examine this possibility.
XRCC1 plays a role in base-excision repair of spontaneous and induced DNA damage. An association of XRCC1 with HPV persistence and CINs was also observed. XRCC1 Arg399Gln (rs25487) also showed an association with differential prognostic factor for progression-free survival in patients with adenocarcinoma of the lung treated with gefitinib chemotherapy [25]. However, in the current study no significant association with cervical cancer risk was observed for polymorphisms in XRCC1 in the BER pathway or NBS1 in the DNA DSBs repair pathway. Our findings suggest that the polymorphisms in ERCC2 and ERCC5 may contribute to the etiology of cervical cancer, showing an additive effect.
High risk HPV infection is prevalent worldwide. However, the actual occurrence of cervical cancer is rare, suggesting the importance of the role of host factors. Rather than a single high penetrance allele, a combination of various genetic factors, each with an individually small effect, is likely to result in predisposition to cervical cancer. More comprehensive screening based on a large number of genetic variants related to cervical cancer susceptibility is expected to provide evidence for addressing cervical cancer-related public health issues such as defining the high risk population for aggressive cancer screening and the use of preventive vaccines.
Our study has a limitation in that only a small number of selected genes and SNPs were examined. In addition, because we had limited epidemiologic information for the patients, we were not able to perform multifactorial combination analysis of the genotype and epidemiological parameters, whereas there was much more information available for the control population. Finally, even though we fully understand the biological significance of the studied genes, conduct of further functional studies will be necessary to illustrate direct effects of these polymorphisms on gene function.
Conclusion
The results of this study show a novel association of cervical cancer and the genes involved in the nucleotide excision pathway in the Korean population. Validation of our results is needed, probably through association studies with sufficient sample sizes, and considering different epidemiological parameters.
Electronic Supplementary Material
Supplementary materials are available at Cancer Research and Treatment website (http://www.e-crt.org).
Notes
Conflict of interest relevant to this article was not reported.
Acknowledgements
We thank M. Maki, K. Sasaki, H. Nagamura, and Y. Morishita for technical support in laboratory analysis.
This research was supported by the International Research & Development Program of the National Research Foundation of Korea (NRF) funded by the Republic of Korea (Grant number: 2013K2A2A4003509, FY 2013), Grant Numbers 1310300 and 1210060 funded by National Cancer Center, Korea, and JSPS KAKENHI Grant Number 13032211-000044.