AbstractPurposeAlthough the incidence of microsatellite instability (MSI) accounts for 10-15% of cases of colorectal cancer, its clinical application for all colorectal cancers has widened. We attempted to identify clinical and pathological parameters that may be helpful in selection of patients with MSI-high (MSI-H).
Materials and MethodsA total of 120 resected colorectal cancers were enrolled retrospectively for this MSI study. Polymerase chain reaction (PCR) and denaturing high performance liquid chromatography and/or real time PCR methods with five markers and immunohistochemistry (IHC) for MLH1 and MSH2 were performed for analysis of cancer and blood specimens. Clinico-pathologic parameters, including IHC, were investigated in order to determine their usefulness as predictive factors of MSI.
ResultsAmong 120 cases of colorectal cancer, MSI was observed in 15 cases (12.5%), including 11 cases of MSI-H and four cases of MSI-low. Patients with MSI were younger, less than 50 years old, had a family history of cancer, Rt. sided colon cancer and/or synchronous multiple colorectal cancer, mucinous histologic type, and serum carcinoembryonic antigen group in the normal range. Results of multivariate analysis showed Bethesda guidelines, Rt. sided and/or synchronous multiple colorectal cancer, and negative expression of IHC for MLH1, which was consistently associated with MSI-H. MSI-H colorectal tumors have met at least one of these three parameters and their sensitivity and specificity were 100% and 72.5%, respectively.
IntroductionMolecular studies have reported that up to 15% of sporadic colorectal cancers and approximately 90% of syndrome of hereditarynon-polyposis colorectal cancer (HNPCC), which accounts for 1 to 3 percent of all colorectal cancers, were caused by inactivation of mismatch repair (MMR) genes, such as MSH2, MLH1, PMS1, PMS2, hMSH6/GTBP, and hMSH3 [1-3]. Inactivation of one of these genes results in DNA microsatellite instability (MSI), characterized by alterations in the length of simple repetitive microsatellite sequences found throughout the genome.While germline mutations of DNA MMR genes have been identified as a causative event in hereditary nonpolyposis, development of sporadic colorectal cancer with MSI-high (MSI-H) is most commonly a result of promoter methylation, leading to epigenetic silencing of the MLH1 gene [4]. Some studies have suggested that colorectal carcinomas with MSI-H tend to have an improved survival rate, and may respond differently to adjuvant chemotherapy than non-MSI tumors [5].
MSI tests may be used routinely as a first-line screening tool for identification of MSI-H colorectal carcinomas for diagnosis of suspected HNPCC and for determination of the clinical implications of MSI-H status in sporadic colorectal cancer. However, to date, MSI tests are complex, time-consuming, and expensive, and are not widely accepted as a screening test.
Clinical data, including family history of cancer, including Amsterdam criteria [6] and/or Bethesda guidelines [7], have been used for selection of suspected HNPCC for MSI testing. In addition to clinical data, a number of studies have suggested that MSI-H colorectal carcinomas may have morphologic characteristics that differ from those of non-MSI-H tumors and therefore may be used as a first-line screening tool for identification of tumors for further molecular testing [8]. However, the utility of histology as a screening tool for MSI-H colorectal carcinomas has yet to be defined [9].
Immunohistochemistry (IHC) for MLH1 and MSH2 protein may increase accuracy of prediction and aid in selection of patients who may have MSI, and it can be used as a screening tool for MSI testing in colorectal cancers [10]. However, these guidelines or histopathologic parameters are not adequate for selection of colorectal cancer patients because their results were studied independently and a false negative result may be obtained.
In this study, we attempted to identify the combination of parameters from clinical data, histopathology, including IHC for MLH1 and MSH2 protein that may be helpful in selection of patients with MSI-H.
Materials and Methods1. Patients and tissue samplesBetween January 2004 and June 2006, the 120 patients who underwent surgery for treatment of colorectal cancers at Daegu Catholic University Hospital, Daegu, Korea were retrospectively enrolled in the present study. Using medical records or telecommunication, the selected patients were investigated successfully for family history of cancer. Detailed family histories were obtained through questionnaires and interviews with patients and their relatives. The questionnaire included cancer history in first- and second-degree relatives and contained questions regarding their age at diagnosis, type of cancer, hospital at which the diagnosis was made, current age, and current status. Patients with HNPCC meeting the Amsterdam criteria, patients with familial adenomatous polyposis, and patients with a vague family history were excluded.
Of the 120 enrolled cases of primary colorectal cancer, 65 were male and 55 were female, with a mean age at the time of surgery of 62.4±11.1 years (range, 36 to 83 years).Twenty patients had a family history of cancer, except hepatocellular carcinoma or cervical cancer, within a second degree pedigree (Table 1). Among them, 10 patients had a family history of colorectal cancer, and only one patient had met the full requirement for diagnosis of Amsterdam, and five were included according to Bethesda guidelines.
The numbers according to tumor location, which were classified as Rt. sided colon (from ileocecal valve to splenic flexure), Lt. sided colon (from descending colon to sigmoid colon), and rectum (from recto-sigmoid colon to distal rectum) and multiple colon (two or more synchronous colon cancers at diagnosis) were 27, 23, 69, and 1, respectively (Table 2).
Tumors in which less than 10% of the cells formed glands were classified as high grade (poorly differentiated), while those containing more than 50% extracellular mucin were classified as mucinous type. The numbers of well/moderately differentiated adenocarcinoma (WD/MD), poorly differentiated adenocarcinoma (PD), and mucinous adenocarcinoma (MU) were 112, 3, and 5, and two were diagnosed as lymphoma.
The depth of invasion to mucosa/submucosa, muscularis layer, pericolic (or perirectal area), or invasion to adjacent other organs were 11, 19, 82, and 6, respectively.
According to International Union against Cancer (UICC) classification, 19 of these tumors were tumor-node-metastasis (TNM) stage I, 55 were stage II, 37 were stage III, and nine were stage IV.
2. Analysis of MSIDNA from blood and tissue were amplified for the five mononucleotide markers of NR21, NR22, NR24, BAT25, and BAT26 [11]. A summary of the amplified locus and primer sequences is shown in Table 3.
Amplification of NR21, 22, and 24 was performed using the home brew PCR system and denaturing high performance liquid chromatography (Transgenomic Inc., Omaha, NE) was used for analysis of the PCR products (Fig. 1).
3. IHC stainOne block of formalin fixed paraffin embedded colorectal cancer tissue was selected for IHC. In all cases, the block was comprised of an area of normal colonic mucosa adjacent to the tumor. Sections (4 µm) were affixed, dried, de-waxed, and rehydrated, followed by inhibition of endogenous peroxidase activity; they were then subjected to heat antigen retrieval. The MLH1 antibody (G168-15, Pharmingen, San Diego, CA) and the MSH2 antibody (G219-1129, Pharmingen) were used for IHC stain using the Streptavidin Biotin Universal Detection System (Immunotech, Marseille, France). Finally, the sections were counterstained in Mayer's haematoxylin.
Among 120 cases, IHC could not be performed for three microsatellite stable (MSS) cases, and, for 117 cases, one pathologist (H.K. Oh), without knowledge of MSI status, performed scoring of IHC staining expression.
4. Statistical analysisFor statistical analysis, the χ2 or Fisher's exact test was used as an appropriate univariate test for selection of optimal clinico-pathologic factors between groups.
We then performed multiple logistic regression for identification of clinico-pathologic factors among all selected optimal factors described above as a univariate test. For selection of clinico-pathologic factors, we used the forward selection method in multiple logistic regression. Using clinico-pathologic parameters identified in multiple logistic regression, we evaluated the prediction of MSI-H as the sensitivity and specificity for the single and all combinations of identified clinico-pathologic parameters. In addition, we used receiver operatring characteristic curve and area under curve to obtain the most appropriate sensitivity and specificity.
The cut off p-value 0.05 was adopted for all statistical analyses. Statistical results were obtained using SPSS ver. 14.0 (SPSS Inc., Chicago, IL).
Results1. Frequency of MSI according to clinical characteristics and carcinoembryonic antigen (CEA) concentrationsOf the 120 tumors, 11 (9.2%) were MSI-H, four (3.3%) were MSI-low (MSI-L), and 105 (87.5%) were MSS (Table 1). Among the 11 patients with MSI-H, there were seven females, which appeared to be higher than that of males, without statistical significance. Ages (mean±standard deviation, years) of MSI-H and MSS patients were 52±9.6 and 63.3±10.9, respectively, which was significantly different (p=0.004). Of patients with a family history of cancer other than hepatocellular carcinoma or cervical cancer, the ratio of MSI-H was 30% (6/20), which was significantly higher, compared to those who had no family history of cancer (p=0.001). Among patients with a family history of colorectal cancer, the ratio of MSI-H was 40% (4/10), which was significantly higher than that of those who had no family history of colorectal cancer (p=0.002). The only patient who met Amsterdam criteria showed MSI-H, and, among five patients who were included according to Bethesda guidelines, three (60%) were MSI-H.
The ratio of increased preoperative serum CEA level was 9% (1/11) in the MSI-H group and 33.3% (33/99) in the MSI-L or MSS group when the cut off value of CEA was regarded as 5 ng/dL (p=0.038).
The ratios of MSI-H in tumors from the right colon and synchronous multiple colon cancers were 29.6% (8/27) and 100% (1/1), which was higher than those of the left colon or rectum (p=0.001) (Table 2).
Among five tumors of MU, three (60%) showed MSI-H, which was higher than those of WD/MD, or PD or lymphoma (p=0.001).
Except for one patient, all tumors with MSI-H invaded to the pericolic (or perirectal) area, and no difference for status of regional lymph node metastasis, lymphovascular invasion, perineural invasion, distant metastasis, or TNM stage was observed between MSI and MSI-L or MSS groups.
2. Positive rates of MLH1 and MSH2 according to MSIThe numbers for expression of MLH1 were 94 (92.2%), 3 (75.0%), and 5 (45.5%), while 8 (7.8%), 1 (25.0%), and 6 (54.5%) were not expressed in MSS, MSI-L, and MSI-H, in order, and the sensitivity and specificity of MLH1 for MSI-H were 54.5% and 92.2%, respectively (Table 4).
In MSH2, the numbers for expression were 101 (99.0%), 4 (100%), and 10 (90.9%), while 1 (1%), 0 (0%), and 1 (9.1%) were not expressed in MSS, MSI-L, and MSI-H, respectively, and the sensitivity and specificity for MSI-H were 9.1% and 99.0%.
3. Identification of parameters using multiple logistic regression analysis for clinico-pathologic and immunohistochemical dataAmong factors that were proved to be statistically significant on a univariate test, multiple logistic regression analysis was performed for identification of parameters that were significantly affected or related to MSI-H.
Immunohistochemical staining for MLH1, tumor location, Bethesda guidelines, and cell types showed a significant association with MSI-H. MSI-H tumors had at least one or more parameters among negative expression of MLH1 protein, right sided colon and/or synchronous multiple colorectal cancer, one of the Bethesda guidelines, or poorly differentiated or mucinous adenocarcinima. Sensitivity and specificity were calculated for all combinations of immunohistochemical staining for MLH1, tumor location, Bethesda guideline, and cell types (Table 5). According to the results, three clinico-pathologic parameters with MLH1, tumor location, and Bethesda guidelines were appropriate and important factors with high sensitivity of 100% and specificity of 72.5%.
DiscussionMSI tests can be applied for diagnosis of HNPCC from all colorectal cancers and for detection of MSI-H tumors in sporadic colorectal cancers. Although a definitive diagnosis of HNPCC could be established by demonstrating a germline mutation, MSI tests can be used as a screening test for suspected HNPCC before mutation analysis. As clinical and prognostic implications in sporadic type colorectal cancer with MSI-H would predict development of multiple synchronous or metachronous-cancers, responsiveness of adjuvant chemotherapy or to chemoprevention related to cyclo-oxygenase-2 inhibitors as well as prognosis, it is also of great importance for identification of MSI-H in sporadic type colorectal cancer in a clinical field [13-15].
For assessment of MSI status, an international consensus meeting held in 1997 proposed a panel of five markers for use in uniform analysis of MSI. This included two mononucleotide repeats (BAT25 and BAT26) and three dinucleotide (D5S346, D2S123, and D17S250) repeats. Tumors showing instability at two or more of these markers were defined as MSI-H, and those showing instability at one repeat or showing no instability were defined as MSI-L and MSS tumors, respectively [7]. In this study, we used five mononucleotide repeats, including BAT25 and BAT26, instead of three dinucleotide (D5S346, D2S123, and D17S250) repeats, because each of the dinucleotide repeats in the aforementioned panel generally show instability in only 60-80% of MSI-H tumors [16]. In addition, when using the Bethesda reference panel, misclassifications of MSI-H tumors occur if two dinucleotide markers were unstable in the absence of BAT26 deletions [17].
As reported by Zhou et al. [18], BAT26 identified the MSI status in 539 of 542 tumors (99.5%), and BAT26 provided the advantage of being a simple and less expensive method that might be used as a screening procedure prior to performance of mutation analysis. According to our data, sensitivity for BAT26 and BAT25 for detection of MSI-H was 100% and 95%, respectively. Our data also indicated the potential of BAT26 for use as a single marker for MSI testing; however, it is not desirable, as there may be the possibility of missing MSI-H by BAT25 or 26 alone due to polymorphisms. BAT25 or BAT26 alone is insufficient for MSI-H screening in all populations [19].
Introduction of the MSI determination as an initial screening test for HNPCC or MSI-H tumors in colorectal cancers has been reported to enable molecular detection of HNPCC in a mass population. Salovaara et al. [20], who examined 535 colorectal cancers, reported that 12% showed MSI and 3.4% had germline mutations of MSH2 or MLH1.This means that at least more than 85% of colorectal cancers are MSS or MSI-L and application of MSI tests for all colorectal cancers may be ineffective due to low sensitivity and specificity for diagnosis of HNPCC or detection of MSI-H in sporadic colorectal cancer.
To save time and cost of MSI testing, some attempts have been made to increase its sensitivity and specificity; these include Bethesda guidelines, evaluation of clinical and histopathologic parameters, MSI tests by one marker or replacement of MSI by IHC of MLH1/MSH2 protein, or a combination of these trials [7-10]. Original Bethesda guidelines for testing colorectal tumors for MSI were proposed in 1997 to guide clinical use to aid in identification of patients who may have HNPCC, and were then revised in 2004 [21]. Bethesda guidelines were established only for diagnosis of HNPCC, not MSI-H sporadic colorectal cancers.
Wullenweber et al. [22] reported that 29% (27/92) of Bethesda-positive patients showed high MSI, compared with 6% (4/72) of patients who did not meet these criteria. According to our data, five patients were included in the Bethesda guidelines and three were MSI-H, including one HNPCC. Use of Bethesda guidelines is a suboptimal method for selection of MSI testsin sporadic colorectal cancer and for application to general medical practice outside academic centers, thus, additional clinical or pathologic data are required for adequate MSI-H tumor selection.
As reported previously, MSI-H tumors are known for having certain clinical and pathological characteristics, including a higher rate of family history of cancer, early age of onset, a tendency for cancers in the proximal colon and excess of poorly differentiated carcinomas or mucinous type, medullary type, contain signet-ring cells, and increased infiltrating T lymphocytes [8]. Ward et al. [23] reported that positive predictabilities of right sided tumor, intraepithelial lymphocytes, poor differentiation, Crohn's-like reaction, mucinous tumor, peritumoural lymphocytes, and Amsterdam criteria for MSI-H in sporadic colorectal cancer were 23%, 35%, 26%, 26%, 20%, 24% and 25%, respectively. Our data also showed similar results, indicating that predictabilities of family history of cancer, Bethesda guidelines, right sided tumor, and poorly differentiation or mucinous tumor for MSI-H in colorectal cancer were 30%, 30%, 28.6% and 14.3%, respectively (data not shown). Alexander et al. [8] reported that medullary carcinoma, intraepithelial lymphocytosis, and poor differentiation were the best discriminators between MSI-H and microsatellite-stable, and suggested that these histopathological evaluations can be used in prioritization of sporadic colon cancers for MSI studies. However, in their study, histopathology alone failed to reliably discriminate MSI-H tumors because the minority of specimens showed no major difference in morphology from the usual MSS cancer. In addition, approximately 40% of MSI-H cancers were not detected, and 6% were never detected by histopathology. These results suggested that these clinical and pathological parameters showed differences between MSI-H and MSS or MSI-L, but are insufficient for clinical application to selection of patients to undergo MSI tests independently. Ward et al. [23] also suggested other parameters associated with MSI-H, including sex, distant metastasis, and stage, however, these findings were not consistent with those of other reports, and our data also did not show any correlation with MSI status.
Engel et al. [24] reported that IHC was highly predictive (99.1%) and specific (99.6%) with regard to MSI tests. However, their data showed that 14 out of 230 (6%) mutations escaped detection by IHC. In our data, the sensitivity and specificity of MLH1 and MSH2 for MSI-H were 54.5%, 92.2% and 9.1%, 99.0%, respectively and among 11 cases of MSIH, seven (63.6%) showed loss of expression of either one of MLH1 or MSH2. and the overall sensitivity and specificity for MSI-H were 63.6% and 92.2%. As reported by Lanza et al. [25], lack of MLH1 nuclear staining was observed much more often than the absence of MSH2 nuclear staining. Despite a close correlation between IHC and MSI, these findings suggested that IHC could not be fully recommended as a substitute for MSI tests.
In our attempt to find parameters showing significant differences between MSS or MSI-L and MSI tumors by multiple logistic regression tests as a multivariate analysis, we found three parameters, as described. Bethesda guidelines, tumors from the right colon and/or synchronous multiple colon cancers, and negative expression of MLH1 protein in MSI-H tumors showed statistical significance. When we applied a selection of patients with one of these three parameters, sensitivity and specificity for MSI-H were 100% and 72.5%, respectively. This means that, using these parameters for selection of patients for MSI testing, we were able to find all MSI-H tumors, with the exclusion of 80 patients who had no MSI-H, and to minimize the number of patients for MSI testing. Using these three parameters, we retrospectively applied 25 patients who underwent surgery before 2003 and were excluded from this study; all five MSI-H patients met at least one of these parameters, and its sensitivity and specificity for MSI-H were 100% and 60%, respectively. These findings allowed us to select patients with MSI-H tumors who met one of three parameters. Except for the Bethesda guideline, IHC, tumor location, and cell type showed 100% sensitivity and 70.6% specificity.
Collection of data for each patient for Bethesda guidelines is not difficult; therefore, careful history taking and routine pathologic data are very important. MSI testing should be recommended for positive patients with any of the Bethesda guidelines, tumor location, or MLH1 stain.
References1. Oh SY, Lee JM, Lee HW, Kim YB, Cheong SH, Kim JM, et al. Clinicopathological significance of microsatellite instability in sporadic colorectal cancer. J Korean Surg Soc. 2006;71:420–425.
2. Sheng JQ, Zhang H, Ji M, Fu L, Mu H, Zhang MZ, et al. Genetic diagnosis strategy of hereditary non-polyposis colorectal cancer. World J Gastroenterol. 2009;15:983–989. PMID: 19248199
3. Woods MO, Younghusband HB, Parfrey PS, Gallinger S, McLaughlin J, Dicks E, et al. The genetic basis of colorectal cancer in a population-based incident cohort with a high rate of familial disease. Gut. 2010;59:1369–1377. PMID: 20682701
4. Wheeler JM, Loukola A, Aaltonen LA, Mortensen NJ, Bodmer WF. The role of hypermethylation of the hMLH1 promoter region in HNPCC versus MSI+ sporadic colorectal cancers. J Med Genet. 2000;37:588–592. PMID: 10922385
5. Watanabe T, Kanazawa T, Tada T, Kazama Y, Hata K, Nagawa H. Chemotherapy and survival in colorectal cancer patients with and without microsatellite instability: can MSI be a prognostic marker? Gastroenterology. 2004;127:688–689. PMID: 15300610
6. Vasen HF, Mecklin JP, Khan PM, Lynch HT. The International Collaborative Group on Hereditary Non-Polyposis Colorectal Cancer (ICG-HNPCC). Dis Colon Rectum. 1991;34:424–425. PMID: 2022152
7. Rodriguez-Bigas MA, Boland CR, Hamilton SR, Henson DE, Jass JR, Khan PM, et al. A National Cancer Institute Workshop on Hereditary Nonpolyposis Colorectal Cancer Syndrome: meeting highlights and Bethesda guidelines. J Natl Cancer Inst. 1997;89:1758–1762. PMID: 9392616
8. Alexander J, Watanabe T, Wu TT, Rashid A, Li S, Hamilton SR. Histopathological identification of colon cancer with microsatellite instability. Am J Pathol. 2001;158:527–535. PMID: 11159189
9. Shia J, Ellis NA, Paty PB, Nash GM, Qin J, Offit K, et al. Value of histopathology in predicting microsatellite instability in hereditary nonpolyposis colorectal cancer and sporadic colorectal cancer. Am J Surg Pathol. 2003;27:1407–1417. PMID: 14576473
10. Shia J, Klimstra DS, Nafa K, Offit K, Guillem JG, Markowitz AJ, et al. Value of immunohistochemical detection of DNA mismatch repair proteins in predicting germline mutation in hereditary colorectal neoplasms. Am J Surg Pathol. 2005;29:96–104. PMID: 15613860
11. Suraweera N, Duval A, Reperant M, Vaury C, Furlan D, Leroy K, et al. Evaluation of tumor microsatellite instability using five quasimonomorphic mononucleotide repeats and pentaplex PCR. Gastroenterology. 2002;123:1804–1811. PMID: 12454837
12. Dietmaier W, Hofstadter F. Detection of microsatellite instability by real time PCR and hybridization probe melting point analysis. Lab Invest. 2001;81:1453–1456. PMID: 11598157
13. Horii A, Han HJ, Shimada M, Yanagisawa A, Kato Y, Ohta H, et al. Frequent replication errors at microsatellite loci in tumors of patients with multiple primary cancers. Cancer Res. 1994;54:3373–3375. PMID: 8012952
14. Karnes WE Jr, Shattuck-Brandt R, Burgart LJ, DuBois RN, Tester DJ, Cunningham JM, et al. Reduced COX-2 protein in colorectal cancer with defective mismatch repair. Cancer Res. 1998;58:5473–5477. PMID: 9850081
15. Carethers JM, Chauhan DP, Fink D, Nebel S, Bresalier RS, Howell SB, et al. Mismatch repair proficiency and in vitro response to 5-fluorouracil. Gastroenterology. 1999;117:123–131. PMID: 10381918
16. Sutter C, Gebert J, Bischoff P, Herfarth C, von Knebel Doeberitz M. Molecular screening of potential HNPCC patients using a multiplex microsatellite PCR system. Mol Cell Probes. 1999;13:157–165. PMID: 10208807
17. Loukola A, Eklin K, Laiho P, Salovaara R, Kristo P, Jarvinen H, et al. Microsatellite marker analysis in screening for hereditary nonpolyposis colorectal cancer (HNPCC). Cancer Res. 2001;61:4545–4549. PMID: 11389088
18. Zhou XP, Hoang JM, Li YJ, Seruca R, Carneiro F, Sobrinho-Simoes M, et al. Determination of the replication error phenotype in human tumors without the requirement for matching normal DNA by analysis of mononucleotide repeat microsatellites. Genes Chromosomes Cancer. 1998;21:101–107. PMID: 9491320
19. Samowitz WS, Slattery ML, Potter JD, Leppert MF. BAT-26 and BAT-40 instability in colorectal adenomas and carcinomas and germline polymorphisms. Am J Pathol. 1999;154:1637–1641. PMID: 10362787
20. Salovaara R, Loukola A, Kristo P, Kaariainen H, Ahtola H, Eskelinen M, et al. Population-based molecular detection of hereditary nonpolyposis colorectal cancer. J Clin Oncol. 2000;18:2193–2200. PMID: 10829038
21. Umar A, Boland CR, Terdiman JP, Syngal S, de la Chapelle A, Ruschoff J, et al. Revised Bethesda Guidelines for hereditary nonpolyposis colorectal cancer (Lynch syndrome) and microsatellite instability. J Natl Cancer Inst. 2004;96:261–268. PMID: 14970275
22. Wullenweber HP, Sutter C, Autschbach F, Willeke F, Kienle P, Benner A, et al. Evaluation of Bethesda guidelines in relation to microsatellite instability. Dis Colon Rectum. 2001;44:1281–1289. PMID: 11584201
23. Ward R, Meagher A, Tomlinson I, O'Connor T, Norrie M, Wu R, et al. Microsatellite instability and the clinicopathological features of sporadic colorectal cancer. Gut. 2001;48:821–829. PMID: 11358903
24. Engel C, Forberg J, Holinski-Feder E, Pagenstecher C, Plaschke J, Kloor M, et al. Novel strategy for optimal sequential application of clinical criteria, immunohistochemistry and microsatellite analysis in the diagnosis of hereditary nonpolyposis colorectal cancer. Int J Cancer. 2006;118:115–122. PMID: 16003745
25. Lanza G, Gafa R, Maestri I, Santini A, Matteuzzi M, Cavazzini L. Immunohistochemical pattern of MLH1/MSH2 expression is related to clinical and pathological features in colorectal adenocarcinomas with microsatellite instability. Mod Pathol. 2002;15:741–749. PMID: 12118112
Fig. 1Microsatellite analysis of NR21 (A), NR22 (B), and NR24 (C) by denaturing high performance liquid chromatography. Microsatellite instability positive specimen (P) showed double peak of chromatogram compared with single peak of control specimen (C). 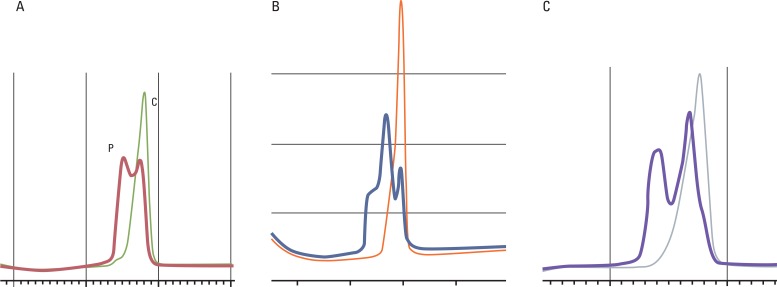
Fig. 2Microsatellite analysis of BAT26 using Light Cycler. BAT26 positive specimens showed shifted melting peaks due to shortening of poly A, as indicated by "P". 
Table 1Clinical characteristics and MSI status Table 2Pathologic characteristics and MSI status Table 3Five mononucleotide markers used for MSI detection Table 4MLH1 and MSH2 expression in colorectal cancers according to MSI status
Table 5Sensitivity and specificity for prediction of MSI-H with single and various combinations of clinico-pathologic characteristics
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||