AbstractCirculating tumor DNA (ctDNA) is the portion of the cell-free DNA in the blood of cancer patients released from tumor cells via apoptosis, necrosis, or active release. From 10 mL of blood, the 4–5 mL of plasma obtained from a cancer patient contains 5–10 ng/mL of ctDNA. The plasma contains not only ctDNA of tumor origin, but also DNA from normal cells or clonal hematopoiesis. Another characteristic of ctDNA is its rapid clearance from circulation; it has a half-life of 16 minutes to 2.5 hours. Obtaining reliable results from ctDNA requires the application and approval of standardized clinical validation guidelines; however, the status of numerous ctDNA tests currently varies. The clinical use of ctDNA testing should be carefully considered based on the test’s specific needs and characteristics. Here we provide the different characteristics of ctDNA tests and information regarding their validation and approval status.
IntroductionAs we currently live in an era of information and advanced genomics, we should maintain our focus on how we respond to and process the overwhelming amount of information we encounter. For example, Mandel and Metais [1] first described the presence of nucleic acids in human blood in 1948, but several decades passed before attention was paid to the vast amount of information supplied by nucleic acids in the blood. However, since the discovery of mutant RAS gene fragments in the blood of cancer patients in 1994 [2] and the detection of microsatellite DNA changes in the serum of cancer patients in 1996 [3], the information contained within the nucleic acids in the blood has gradually gained attention.
Blood contains cellular components and numerous biological substances, such as extracellular vesicles, proteins, and nucleic acids, including mRNAs, miRNAs, and cell-free DNA (cfDNA). cfDNA refers to any non-encapsulated DNA within the bloodstream originating from various cell types. The portion of the cfDNA in the blood of cancer patients released from tumor cells via apoptosis, necrosis, or active release [4,5], is commonly referred to as the circulating tumor DNA (ctDNA). The ctDNA has gained increasing attention since 2010 because of the potential to detect early cancer metastases through novel, sensitive laboratory methods that cannot be detected by high-resolution imaging techniques [6]. The BRACAnalysis (Myriad Genetic Laboratories, Salt Lake City, UT) was the first Food and Drug Administration (FDA) approved companion diagnostic test using ovarian cancer patient’s blood specimens with the development of a gene mutation treatment. Expectations arose that ctDNA could lead to drug treatments for cancer patients. Since then, the number of tests and studies related to ctDNA has exploded exponentially (Fig. 1).
Nucleic acids in the blood are heterogenous depending on their origin. ctDNA analysis can provide more comprehensive information than a conventional tissue biopsy, which has the spatial limitation inherent in sampling due to tumor tissue heterogeneity. It is estimated that up to 3.3% of tumor DNA enters the blood daily from 100 g of tumor tissue, equivalent to 3×1010 tumor cells [7]. On average, the size of the ctDNA varies from small fragments of 70–200 base pairs to large fragments of up to 21 kb [8]. It is important to note the relatively short half-life of ctDNA in blood circulation, ranging from 16 minutes to 2.5 hours [9,10]. Although many tumor-specific abnormalities (e.g., mutations in tumor or tumor suppressor genes, changes in DNA integrity [11], abnormal gene methylation, changes in microsatellite [3], mitochondrial DNA loading levels, and changes in chromosomal genomes [12]) can be detected using ctDNA, a number of obstacles exist in the implementation of ctDNA for screening and diagnosis. First, normal hematopoietic cells and other nucleic acids of non-tumor origin also contribute to the ctDNA in the blood and cause false positives in ctDNA assays in cancer patients [13,14]. Not all somatic mutations detected in the ctDNA analyses are of cancer origin; clonal expansion of somatic variants can be observed in healthy individuals and may represent clonal hematopoiesis of indeterminate potential (CHIP). CHIP frequency increases with age, with only 1% of people under the age of 50 but > 10% over the age of 65 exhibiting CHIP [15–17]. These abnormalities commonly occur in the DNMT3A, TET2, and ASXL1 genes [16], but have also been reported in other genes such as TP53, JAK2, SF3B1, GNB1, PPM1D, GNAS, and BCORL1 [15]. Simultaneous occurrence of CHIP and tumor-derived gene mutations that have abnormalities in these genes may cause difficulties interpreting the ctDNA assays. The second issue is the low concentration of ctDNA (1–10 ng/mL in asymptomatic individuals) [18]. Depending on the concentration of ctDNA, a false negative result is possible; therefore, the sample volume is an important factor affecting the results. Third, the variant allele frequency (VAF) of ctDNA is usually much lower, often below 1%, and can be affected by factors such as cancer type, stage, and clearance rate [19]. Any interpretation of the results requires careful decisions regarding the threshold of allele frequencies of the detected variants, as these are critical aspects. Fourth, there is a lack of consensus on how ctDNA detection should be performed, from the extraction stage to the final in silico variant analysis stage. Even the nomenclatures related to ctDNA lack a proper consensus [20]. Due to the rapid incremental clinical use of ctDNA testing, the unmet demand for a proper consensus on ctDNA-related issues remains [20,21].
This review describes the currently-available ctDNA assays based on the different methodologies, ranging from the traditional methods to more recent advanced molecular technologies. We focus on the unmet need for clinical validation of ctDNA testing by reviewing the validation and approval processes of the FDA and European Commission in vitro Diagnostic Medical Device (CE-IVD), among others. This review addresses frequently raised questions regarding the clinical application of ctDNA assays, summarizes the current status of approved and validated ctDNA assays, and the future direction of ctDNA testing.
The A to Z of the ctDNA TestBefore introducing the ctDNA test, the terminology and definitions of ctDNA must be clarified. Bronkhorst et al. [22] proposed a nomenclature system for three highly investigated diagnostic areas based on the biological compartment in which the cfDNA is distributed (depending on its presence in circulation) and the origin (Fig. 2) [23]. cfDNA is highly heterologous, and a broader concept is needed that covers both nuclear and microbial DNA. Nuclear DNA includes mitochondrial DNA, and microbial DNA encompasses both microbial and viral DNA, not of human origin. Part of the nuclear DNA in the plasma of cancer patients is ctDNA. ctDNA usually refers to all types of tumor-derived DNA in the circulating blood, as discussed in this review.
DNA abnormalities occur by different parts, and each has different features. These features of the ctDNA have different potential clinical implications (Fig. 3). Genomic aberrations of somatic origin detectable in the ctDNA include mutations, chromosomal rearrangements, and copy number changes. An additional features characteristic of ctDNA is specific epigenetic aberrations such as methylation patterns or different DNA fragment lengths [24]. Although ctDNA tests vary in their genomic features and coverage of the genes of interest, the basic principles of the test remain the same. Two categories exist; targeted approaches that test for a small number of known mutations, and untargeted approaches that broadly test for unknown targets. A targeted approach includes real-time polymerase chain reaction (RT-PCR), digital PCR (dPCR), and beads, emulsion, amplification, and magnetics (BEAMing) technology, whereas the broader approach would include high-throughput sequencing methods based on next-generation sequencing (NGS), whole exome sequencing (WES), whole genome sequencing (WGS), and mass-spectrometry-based detection of PCR amplicons [25] among others.
1. ctDNA detection methods1) RT-PCRRT-PCR is widely used for variant screening because it is relatively inexpensive and fast [26]. The variants are detected via the binding of complementary sequences using fluorescent-labeled sequence-specific probes, and the fluorescence intensity is related to the amount of amplified product. The sensitivity of RT-PCR is approximately 10%, which is lower than that of other test methods [27,28]. Cold amplification at a lower denaturation temperature PCR (COLD-PCR) is a variant assay that improves the RT-PCR sensitivity. COLD-PCR concentrates mutated DNA sequences in preference to the wild type using a lower-temperature denaturation step during the cycling protocol. The denaturation temperature for a given sequence is adjusted within ±0.3°C to allow for selective denaturation and amplification of mutated sequences, while double-stranded wild-type sequences are amplified less. This assay can enrich the mutant sequences, improving the sensitivity to detect the mutant allele frequency (MAF) to approximately 0.1% [29,30]. The PCR-based method has the advantage of high sensitivity and cost-effectiveness but is limited as only known variants can be selected with limiting input and speed.
2) Digital PCRdPCR shares the same reaction principle as RT-PCR, except that the samples are dispersed into arrays or droplets, resulting in thousands of parallel PCR reactions. The dPCR can quantify a low fraction of variants against a high background wild-type cfDNA using a single or few DNA templates in an array/droplet and has 0.1% sensitivity [26]. The dPCR method can be applied to cancer personalized profiling by deep sequencing (CAPP-Seq) in combination with molecular barcoding technologies that improve sensitivity by reducing the background sequencing error [31,32]. These two methods improve the sensitivity of CAPP-Seq up to three-fold and, when combined with molecular barcoding, yield approximately 15-fold improvements [32]. cfDNA enrichment is conducted by a two-step PCR procedure during the sample preparation process. The first PCR amplifies the mutational hotspot regions of several genes in a single tube. The second PCR is a nested PCR with unique barcoded primers for sample labeling. The final PCR products are pooled and partitioned for sequencing. The advanced dPCR assay (BEAMing) is a highly sensitive approach with a detection rate of 0.02% [33,34]. This approach consists of four principal components: beads, emulsion, amplification, and magnetics. BEAMing combines dPCR with magnetic bead and flow cytometry [27,33]. In BEAMing, the primer binds to the magnetic beads using a reaction that forms a biotin-streptavidin complex. Less than one template molecule and less than one bead are contained within the microemulsion, and the PCR is performed within each droplet. At the end of the PCR process, the beads are magnetically purified. After denaturation, the beads are incubated with oligonucleotides to distinguish between different templates. The bound hybridization probe is then labeled with a fluorescently labeled antibody. Finally, the amplified products are counted as fluorescent beads by flow cytometry. However, the BEAMing method is impractical for routine clinical use due to its workflow complexity and high cost [34,35].
3) Mass spectrometryThe mass spectrometry-based method combines the matrix-assisted laser desorption/ionization time-of-flight mass spectrometry with a conventional multiplex PCR. An example of this method is UltraSEEK (Agena Bioscience, San Diego, CA). UltraSEEK consists of two-step PCR for amplification and mass spectrometry for detection. The two-step PCR step consists of a multiplex PCR followed by a mutation-specific single-base extension reaction. The extension reaction uses a single mutation-specific chain terminator labeled with a moiety for solid phase capture. Captured, washed, and eluted products are examined for mass, and mutational genotypes are identified and characterized using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry [36]. UltraSEEK has the advantage of multiplex detection of mutant sequences simultaneously, and has a MAF of 0.1% [37].
4) Next-generation sequencingNGS, also known as massively parallel sequencing technology, can characterize cancer at the genomic, transcriptomic, and epigenetic levels. NGS is a highly sensitive assay that can detect mutations in MAF of < 1% using the latest platforms [38]. NGS can analyze several million short DNA sequences in parallel and conduct sequence alignment or de novo sequence assembly to the reference genome [39]. Depending on the panel configuration, NGS panels can be targeted to analyze known variants or untargeted to screen unknown variants. Target panels were preferred due to their high sensitivity and low cost but are limited to point mutations and indel analysis. Several NGS methods can be applied to target panels with adjustable sensitivity, including tagged amplicon deep sequencing, the safe sequencing system, and CAPP-Seq. On the other hand, WGS or WES using untargeted panels allow detection of unknown DNA variants throughout the entire genome (or exome). Different genome-wide sequencing methods have been proposed for different variation types, such as personalized analysis of rearranged ends, digital karyotyping, and Fast Aneuploidy Screening Test-Sequencing System [35]. However, genome-wide sequencing requires a large sample, making its application for ctDNA difficult due to the low concentrations of ctDNA in samples.
There have been attempts to analyze DNA fragmentation differences. There is a marked difference in fragment length size between ctDNA and normal cf DNA. The fragment length of ctDNA is consistently shorter than that of normal cfDNA [40]. Besides, ctDNA with a low MAF (< 0.6%) is associated with a longer ctDNA fragment length when compared to normal cfDNAs [41]. Moreover, most cancers of different origins showed fragmentation profiles of varying lengths [42]. The characteristic DNA fragmentation provides a proof-of-principle approach applicable to screening, early detection, and monitoring of various cancer types.
NGS application has been extended to microsatellite inst-ability (MSI) detection [43]. Loss of DNA mismatch repair (MMR) activity leads to an accumulation of mutations that could otherwise be corrected by MMR genes. A deficiency in MMR activity is often caused by germline mutations or aberrant methylation. The MSI phenotype of a deficiency in MMR activity refers to the shortening or lengthening of tandem DNA repeats in coding and noncoding regions throughout the genome. Tumors with at least 30% to 40% of unstable microsatellite loci, termed microsatellite instability high (MSI-H) [43], reportedly have a better prognosis than MSS tumors and tumors with low MSI. MSI has been documented in various cancer types, including colon, endometrium, and stomach cancers [44,45]. The FDA has approved Pembrolizumab to treat MSI-H cancer regardless of the tumor type or site [46]. NGS-based methods utilize various MSI detection algorithms such as MSIsensor [47], mSINGS [48], MANTIS [49], and bMSISEA [43], which have demonstrated concordance rates ranging from 92.3% to 100% with the PCR-based method. The application of NGS can reliably detect the MSI status with a ctDNA fraction up to 0.4% [43].
5) Methylation analysisEpigenetic information such as methylation is more specific to the tissue of origin than genetic mutations [50]. Changes in DNA methylation patterns occur early in tumor development and have been reported to help early screening for cancers of unknown origin [35,51]. Methylation analysis is not routinely or commonly used to detect ctDNA, but it can be partially applied to cancer patients. The method can be broadly divided depending on its application to the candidate gene. The Grail’s technology applied DNA methylation patterns to differentiate between cancer cell types or tissue origins [52]. Most cfDNA methylation analysis methods applied a candidate gene approach due to the low analytical cost and the efficiency of using pre-established epigenetic biomarkers [53]. Bisulfite treatment-based assays distinguish cytosine methylation and are generally the preferred ctDNA methylation detection method [54]. The analytical principle is based on treating the DNA with bisulfite to convert unmethylated cytosine residues to uracil. Two types of ctDNA methylation analysis exist; PCR-based methods that apply specific primers or melting temperatures and sequence-based methods such as direct sequencing or pyrosequencing. However, the accuracy of bisulfite pyrosequencing is only maintained up to 5% [55,56]. Methylation-specific PCR (MSP) can distinguish DNA sequences by sequence-specific PCR primers after bisulfite conversion [57]. The methylation-sensitive high-resolution melting (MS-HRM) protocol is based on comparing the melting profiles of the PCR products from unknown samples with profiles for specific PCR products derived from methylated and unmethylated control DNAs [58]. The protocol consists of PCR amplification of bisulfite-modified DNA with primers and subsequent high-resolution melting analysis of the PCR product. MSP or MS-HRM can accurately detect about 0.1% of methylated DNA [57,58].
6) Hybrid sequencing (NanoString)The nCounter Technology (NanoString Technologies, Seattle, WA) is a novel technology developed to screen clinically-relevant ALK, ROS1, and RET fusion genes in lung cancer tissue samples. NanoString is applicable to RNA, miRNA, or protein and, more recently, to ctDNA [59–61]. Target ctDNA is directly tagged with capture and reporter probes that are specific to the target variant of interest, creating a unique target-probe complex. The probes include a fluorescent reporter and a secondary biotinylated capture probe that allows immobilization onto the cartridge surface. The target-probe complex is immobilized and aligned on the imaging surface. The labeled barcode of the complex is then directly counted by an automated fluorescence microscope [62,63].
2. International efforts for advanced precision medicine in ctDNA analysisThe availability of new ctDNA testing methods and continuous scientific advances has resulted in several new problems. The factors affecting ctDNA testing outcomes are present from the sample collection phase to the final reporting phase. The American Society of Clinical Oncology (ASCO) and the College of American Pathologists (CAP) reviewed the framework for future research into clinical ctDNA tests in 2018 [64]. This article categorizes the key findings that affect ctDNA testing for oncology patients into preanalytical variables for ctDNA specimens, analytical validity, interpretation and reporting, and clinical validity and utility of each test. Plasma is the most suitable sample recommended for ctDNA testing [65], as is the use of specific types of sample collection tubes such as cell-stabilizing tubes (Cell-Free DNABCT [STRECK tubes] and PAXgene Blood DNA tubes [Qiagen]), or conventional EDTA anticoagulant tubes [66–68]. Leukocyte stabilization tubes can extend the preprocessing window to 48 hours after collection, but EDTA anticoagulant tubes require processing within 6 hours. However, few studies have examined the preanalytical variables affecting ctDNA testing, and guidelines are needed to validate their clinical utility. Considering the variations in the many factors and different types of ctDNA assays, based on different methods, the validity of each analysis must be comparable. The current clinical ctDNA analyses require a clear assessment of the validity of the individual analyses. To increase the precision of ctDNA assays, best practices, protocols, and quality metrics for NGS-based ctDNA analyses must be developed.
The Sequencing Quality Control Phase 2 (SEQC2) consortium organized by the FDA is an international group of members from academia, government, and industry (https://www.fda.gov/science-research/bioinformatics-tools/microarraysequencing-quality-control-maqcseqc#MAQC_IV). The SEQC2 Oncopanel Sequencing Working Group developed a translational scientific infrastructure to be applied for practices in precision oncology [69]. The Oncopanel Sequencing Working Group evaluated panels/assays, genomic regions, coverage, VAF ranges, and bioinformatics pipelines, using self-constructed reference samples. This study on the analytical performance evaluation of oncopanels/assays for small variant detection includes: (1) comprehensive solid tumor oncopanel examination [70], (2) liquid biopsy testing [71], (3) testing involving formalin-fixed paraffin-embedded material [72], and (4) testing involving spike-in materials [73]. A major finding from the SEQC2 liquid biopsy proficiency testing study [71] is that all assays could detect mutations with high sensitivity, precision, and reproducibility for those above the 0.5% VAF threshold. The degree of DNA input material impacted the test sensitivity, requiring higher input for improved sensitivity and reproducibility for variants with a VAF below 0.5%.
Advanced NGS-based assays for precision oncology are in high demand, and recently approved ctDNA assays are to be identified. Establishing a proper validation scheme would support the FDA’s regulatory and scientific endeavors.
3. FDA-approved ctDNA assayWe searched for FDA-approved assays in the FDA database (https://www.accessdata.fda.gov/scripts/cdrh/devicesatfda/index.cfm) using the following keywords: circulating tumor DNA, ctDNA, cell-free DNA, circulating cell-free DNA, cfDNA, liquid biopsy, and plasma and DNA. The search results were compared to the annual report of medical devices cleared or approved on FDA lists published between 2013 and 2022 for confirmation, and assays related to ctDNA were selected. We identified three in vitro diagnostic devices (Epi ProColon, Cobas EGFR Mutation Test, and therascreen PIK3CA RGQ PCR Kit) and two specialized laboratory services (Guardant360 CDx, and FoundationOne Liquid CDx) (Fig. 4).
FoundationOne Liquid CDx was approved as a companion diagnostic on October 26 and November 6, 2020. The approved companion diagnostic indications are (1) to identify mutations in BRCA1 and BRCA2 genes in patients with ovarian cancer eligible for treatment with rucaparib (RUBRACA, Clovis Oncology, Inc.), (2) to identify ALK rearrangements in patients with non–small cell lung cancer eligible for treatment with alectinib (ALECENSA, Genentech USA Inc.), (3) to identify mutations in the PIK3CA gene in patients with breast cancer eligible for treatment with alpelisib (PIQRAY, Novartis Pharmaceutical Corporation), and (4) to identify mutations in the BRCA1, BRCA2, and ATM genes in patients with metastatic castration-resistant prostate cancer eligible for treatment with olaparib (LYNPARZA, AstraZeneca Pharmaceuticals LP) [74]. The NGS-based ctDNA tests related to companion diagnostics, such as FoundationOne Liquid CDx and Guardant360 CDx are transitioning to specialized laboratory services. FDA-approved tests require evaluations of their analytical performance. Recent laboratory-based tests have undergone extensive evaluation testing using large sample numbers for advanced assay interpretation and reporting, clinical validation, and utility (Table 1). Considering the cost of the tests, the number of tests performed for evaluation is prohibitive for small laboratories. Specialized laboratories use their own processes (Table 2) to provide users with reports. Therefore, testing is changing from a complex assay performed at individual laboratories to a more specialized service where each specialized laboratory be devised its own analysis processes, and the type of inspection changes to a laboratory service.
4. CE-marked ctDNA assayIn May 2017, the Conformité Européenne (CE) declared the strengthening of in vitro Diagnostic Regulation and Medical Device Regulation regulations. The transition was completed in May 2022, following a 5-year transition period. The CE announced a new database search service called the European Database on Medical Devices (EUDAMED), which will consist of six modules: actor registration, unique device ID and device registration, notification authority and certificate, clinical and performance research, and alert and market monitoring. The EUDAMED database was scheduled for release in July 2022 but has been postponed to the third quarter of 2024 due to a delay in the module development process. We had difficulty performing a systematic search for medical devices or in vitro diagnostics with the CE mark. Therefore, we searched for recently published papers that mentioned CE-marked products (Table 3).
5. Marketplace of ctDNA test in Republic of KoreaIn Korea, the marketplace of ctDNA test has recently expanded as the In Vitro Diagnostic Medical Devices Act was promulgated in April 2019. When ctDNA test was searched in the medical device database of the Korea Ministry of Food and Drug Safety (https://udiportal.mfds.go.kr/search/data/P02_01#list), a total of seven domestic tests were identified. The Smart Biopsy EML4-ALK Detection Kit (CytoGen, Seoul, Korea) was first nationally accredited on April 20, 2016, and the following tests have been nationally accredited in sequence; ADPS EGFR Mutation Test Kit V1 (GENECAST, Seoul, Korea), Droplex KRAS Mutation Test v2 (Gencurix, Seoul, Korea), Droplex PIK3CA Mutation Test (Gencurix), PANAMutyper R EGFR V2 (PANAGENE, Daejeon, Korea), AlphaLiquid 100 (IMBDX, Seoul, Korea), and LiquidSCAN (GENINUS, Seoul, Korea). Most tests are PCR-based assay, such as RT-PCR and dPCR, but AlphaLiquid 100 (IMBDX) and LiquidSCAN (GENINUS) are NGS-based assay.
Current and Future DirectionsThe introduction of ctDNA testing and technical advancements in NGS have affected cancer’s diagnostic and therapeutic aspects. Many biomarkers associated with treatment options have been identified for cancer patients whose tissues were previously unavailable for biopsies. The widespread use of NGS and its increased availability has changed the concept/scheme of companion diagnostics from ‘one gene-one drug’ to ‘multi-genes - multi-drugs’ treatment [75]. Recently, experts from the National Comprehensive Cancer Network have recommended measuring multiple predictive genes associated with companion diagnostics for certain cancers [76].
Although tissue biopsy remains the standard of diagnosis because of its important pathological diagnostic information value and the need to assess biomarkers without DNA alterations, such as estrogen receptor expression and other protein or RNA biomarkers [77]. However, ctDNA testing is undoubtedly a very promising technology, with broad clinical applications for early diagnosis, monitoring, management, and prognosis [78,79]. When performing metastatic diagnosis alongside standard tissue biopsies, ctDNA testing can provide key advantages, either as a baseline for follow-up testing after treatment or in situations in which more-rapid identification of targetable alterations is needed to guide first-line therapy [77]. In addition, ctDNA testing plays an important role in real-time monitoring of various aspects of tumors due to its simple sample preparation [78].
We expect that the strengths of ctDNA, including the potential ability to detect latent cancers and track tumor-specific mutations, will naturally enable minimal residual disease (MRD) assessment [80]. The ability to identify microscopic residuals and occult metastases could revolutionize the individualization of adjuvant and consolidation therapy [81]. Despite the potential use of ctDNA to determine MRD, it is premature for use in this feature due to many current issues [82,83]. Therefore, the reliability and clinical validity of ctDNA analysis is becoming increasingly important as it can directly impact patient care with respect to treatment options.
To assess the current status of ctDNA testing and ongoing developments, we searched the clinical trial database of the FDA (Clinical trial.gov). A query using ctDNA as the keyword showed 978 clinical trials as of June 2022. The results of 109 trials in which the clinical trials were completed were reviewed using the uploaded articles from the database or the national clinical trial number for a PubMed search. Twenty-two clinical trials were available, and we reviewed and compared their preanalytical and analytical variables (Fig. 5). The preanalytical variables of the blood collection tube used, the volume of whole blood collected, time to sample processing, centrifugation protocols, and DNA extraction methods were missing or unidentifiable in over half of the reports, despite their importance. When provided, the information varied among studies; specimen processing within 24 hours using EDTA tubes was a possible confounding factor regarding the stability of the ctDNA. Some trials requiring detection of low VAF variants, such as ‘using copy number variation of ctDNA for cancer diagnosis’ or ‘biomarker response according to treatment in metastatic cancer’ used only 25 ng DNA, which appears to be insufficient and, therefore, they were unable to exclude the possibility of false negatives. The use of FDA-approved assays among trials was low at 13.64% (3/22). Information regarding approval from other institutions or agencies was often unavailable, but most clinical trials (72.73%, 16/22) utilized non–FDA-approved testing methods.
Despite the considerable therapeutic influence of ctDNA testing or companion diagnostics, the current practice of utilizing various ctDNA tests without regard to a consensus on clinical validation is questionable, as the review of clinical trials and available information demonstrates. Discrepant test results between the tissue biopsy and ctDNA results are common, and the underlying reasons for these discrepancies include temporal heterogeneity (an archival tumor specimen), spatial heterogeneity (a subclonal mutation), and analytical errors [64]. In the case of analytical errors, the source of the error should be evaluated before any therapeutic action can be taken. If such an investigation or validation is lacking, this should be disclosed to enable the participants or patients to give proper informed consent. The common rules followed by institutional review boards (IRB) when reviewing research state that the prospective participants (or a legally authorized representative) be provided with sufficient detailed information regarding the research. The consent form containing this information must be organized to facilitate an understanding of why one might or might not want to participate [84,85]. This should also be the case if the patient opts for a ctDNA test. Patients should be able to choose the ctDNA test based on detailed information about the accuracy of the test, the list of genes that can be analyzed, and the laboratory’s ability to analyze mutations based on experience.
Laboratories must be aware of any new developments in ctDNA testing. A changing trend in ctDNA testing is demonstrated by the recent FDA-approved ctDNA assays. Previously FDA-approved assays were mostly in vitro diagnostic devices (IVDs), conducted by small-scale clinical laboratories. However, recent FDA-approved assays require referrals to larger, specialized laboratories with institutional accreditation. Such changes are inevitable due to the testing complexity and higher reliability required by clinical practice. Cutting-edge ctDNA testing is costly and requires first-rate laboratory infrastructure and highly specialized and multi-disciplinary professionals. The trend toward centralization and referrals is in line with these requirements.
Previously, the introduction of tumor markers has resulted in the overutilization of tumor marker testing in the hope of providing definitive answers to cancer diagnostics. The need for specific guidelines/instructions on how tumor markers should be utilized demonstrates the concern regarding the misuse of tumor markers, potentially resulting in misdiagnosis or a delay in treatment [86]. In recognition of these issues, in 2002, the National Academy of Clinical Biochemistry produced the Laboratory Medicine Practice Guideline of tumor biomarkers [87]. The guideline provides recommendations based on expert opinions from those in the field of IVD and the marketplace. This regulatory guideline includes 16 different cancers and their established tumor markers, their qualities, and the technological requirements. It is anticipated that a similar development to the guideline for tumor marker testing will be available for ctDNA testing soon. However, guidelines on validation are currently lacking. ctDNA testing requires clinical validation prior to its clinical implementation. These regulated clinical validation guidelines will inevitably require updating, refinement, and modification as knowledge and understanding of ctDNA and its biological role increases.
In summary, ctDNA testing requires a minimum safety resolution through clinical validation to ensure its clinical utility. The testing requires cooperation between multi-disciplinary experts to provide meaningful and reliable results. Establishing a proper clinical validation guideline for ctDNA will enable access to better cancer treatment and reliable testing in the future.
NotesFig. 1The number of publications searched on the PubMed database (https://pubmed.ncbi.nlm.nih.gov/) from 1974 to 2021 using the following keywords: cell-free DNA, cfDNA; circulating tumor DNA, ctDNA; and liquid biopsy. Search was limited to exact match for each term, and the species limited to human. The liquid biopsy was first reported in 1974, and cell-free DNA was first reported in 1986. Since 2014, the number of publications for all search terms, including circulating tumor DNA has increased exponentially. The rapid increase in publications resulted in fragmented use of similar nomenclatures, and the number of search results using terms with the same meaning are displayed differently. 
Fig. 2Defining a systematic nomenclature for confounding terms regarding circulating tumor DNA (ctDNA). The ctDNA is a concept that belongs to cell-free nucleic acid in a broad sense and cell-free DNA in a narrow sense. The cell-free nucleic acids are biologically and structurally diverse. Depending on the origin of the nucleic acids, the nucleic acids are mixed, such as nucleic acids derived from various cells or microorganisms. For systematic nomenclature, a top-down approach from cell-free nucleic acid into subclasses are necessary. First, nucleic acids can be classified according to its presence in circulation such as blood and lymphatics, or in non-circulatory fluids, such as urine, saliva, and cerebrospinal fluid. As the use of term cell-free nucleic acids is not limited to cancer, but also applied in prenatal testing and post-transplant surveillance, the terms should be organized by their classifications. Figures were created with BioRender [23]. 
Fig. 3Different features of circulating tumor DNA (ctDNA) in the plasma. Different tumor-related clinical information can be obtained depending on the structural features of the ctDNA detected during the examination. In ctDNA, DNA abnormalities related to cancer are identified as somatic mutations, copy number aberrations, and structural abnormalities of chromosomes such as inversions, translocations, insertions, and deletions. In addition, useful information can be obtained through epigenetic aberrations such as methylation patterns and DNA fragment size distributions. Figures were created with BioRender [23]. 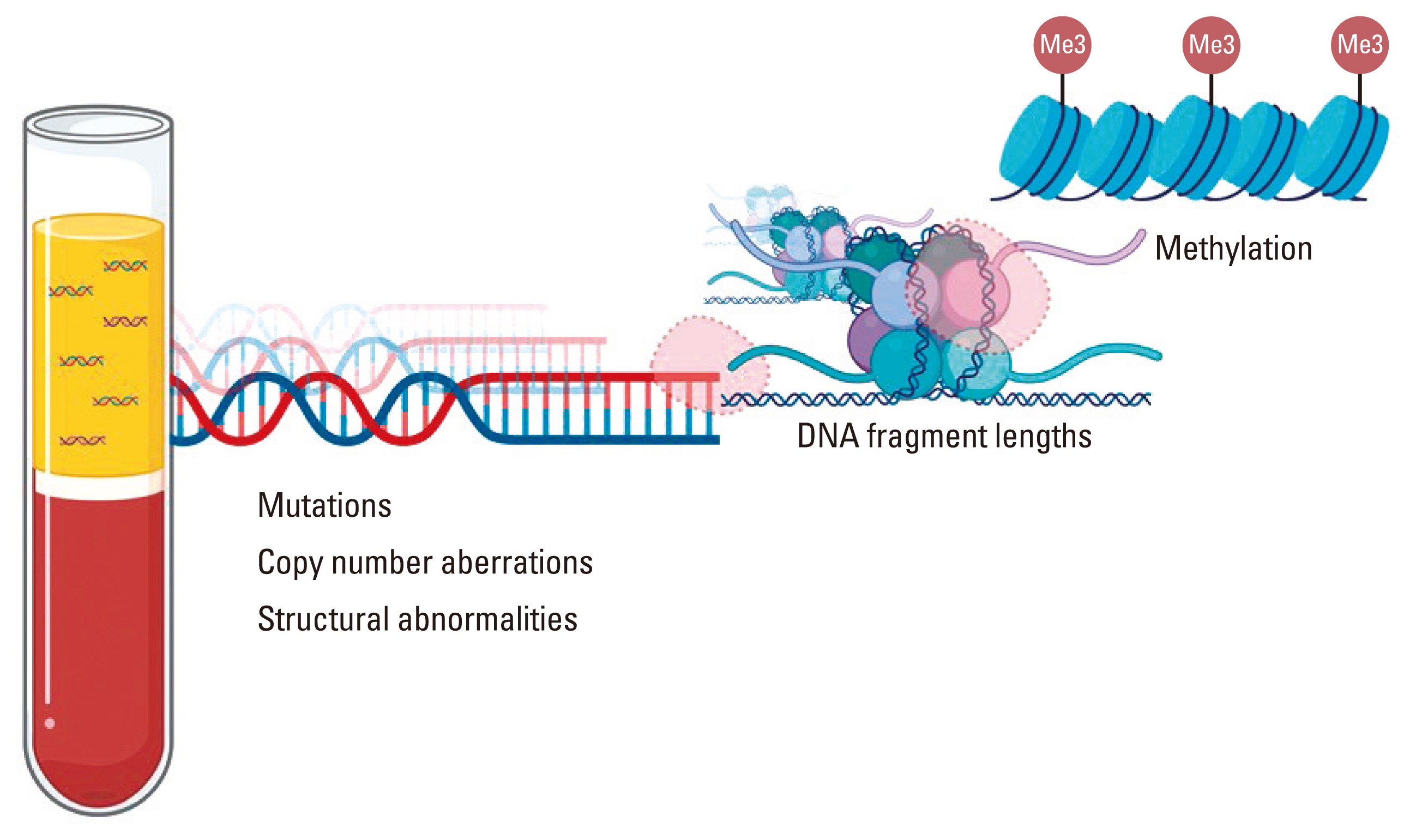
Fig. 4A brief summary of the FDA-approved assays in chronological order. Assays are presented in the order of FDA approval date (circle) placed within the timeline (red line). The assays listed above are categorized as in vitro diagnostic devices and assays provided as laboratory services are listed under the timeline. ALK, anaplastic lymphoma kinase; AS-PCR, allele-specific PCR; BCT, blood collection tube; EGFR, epidermal growth factor receptor; cfDNA, cell-free DNA; FDA, Food and Drug Administration; FFPE, formalin-fixed paraffin-embedded; N/A, not applicable; NGS, next-generation sequencing; RT-PCR, real time-polymerase chain reaction. 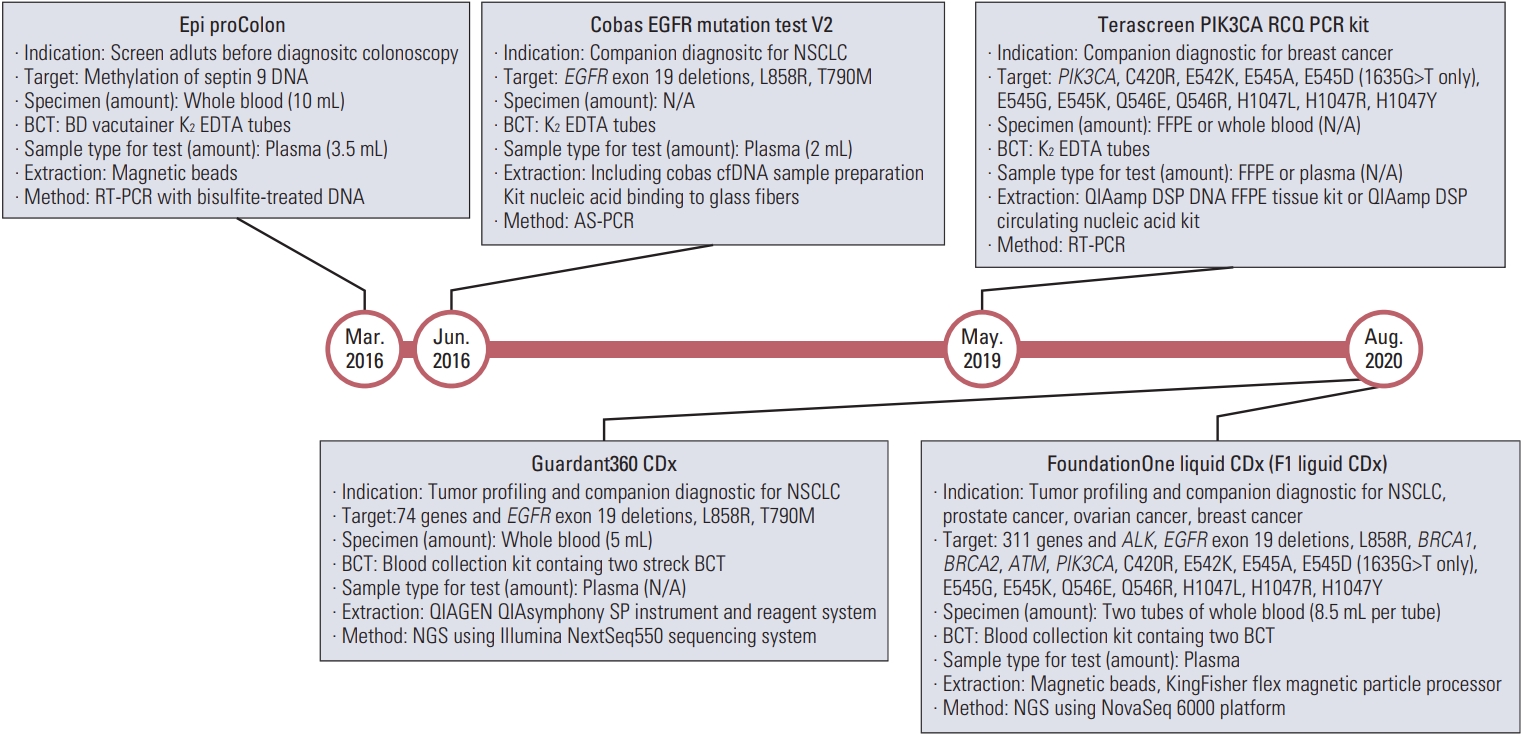
Fig. 5Current variances present among circulating tumor DNA (ctDNA) testing used in clinical trials. The following critical testing conditions were identified from the clinical trials using ctDNA provided from ClinicalTrials.gov when available; blood collection tube (A), whole blood volume (B), time to sample processing (C), 1st centrifugations (D), 2nd centrifugations (E), DNA extraction method (F), input DNA volume (G), and use of Food and Drug Administration (FDA)–approved test (H). The degree of agreement in factors of whole blood volume, sample processing methods used for ctDNA acquisition, and utilization of FDA-approved assay was low. 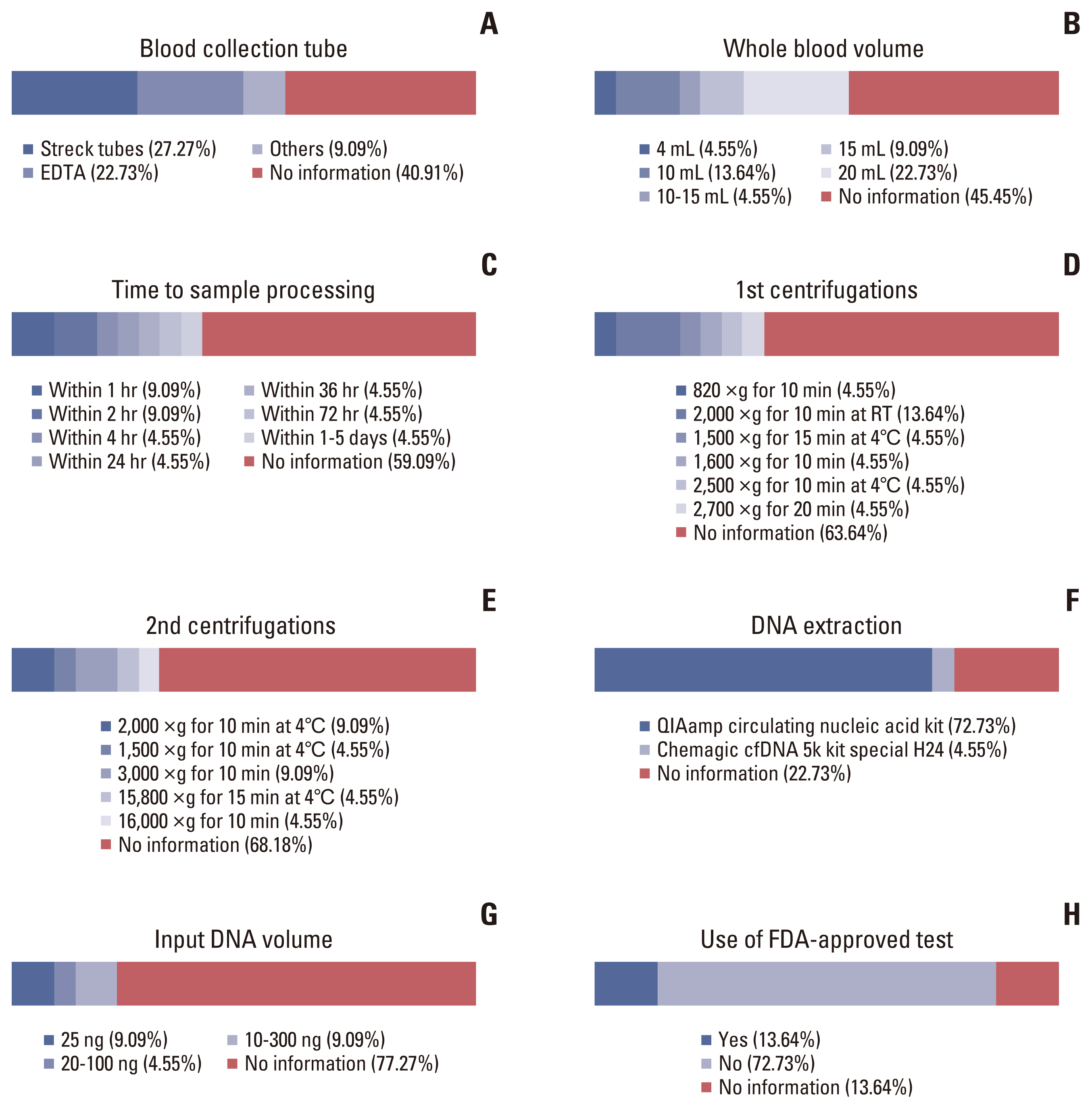
Table 1Total number of tests performed in validation studies for FDA-approved ctDNA tests
AS-PCR, allele-specific PCR; cfDNA, cell-free DNA; ctDNA, circulating tumor DNA; EGFR, epidermal growth factor receptor; FDA, Food and Drug Administration; IVD, in vitro diagnostics device; LDT, laboratory-developed tests; LoB, limit of blank; LoD, limit of detection; NGS, next-generation sequencing; RT-PCR, real-time reverse transcription polymerase chain reaction. a) See FDA approval documentation (https://www.accessdata.fda.gov/scripts/cdrh/devicesatfda/index.cfm). Table 2Variables for interpretation of NGS-based FDA-approved ctDNA tests
CNA, copy number alterations; ctDNA, circulating tumor DNA; FDA, Food and Drug Administration; Indel, small insertion/deletions; MAF, mutant allele frequency; NGS, next-generation sequencing; SNV, single nucleotide variants. a) See FDA approval documentation (https://www.accessdata.fda.gov/scripts/cdrh/devicesatfda/index.cfm). Table 3The list of CE-IVD and FDA-approved ctDNA tests
References1. Mandel P, Metais P. Nuclear acids in human blood plasma. C R Seances Soc Biol Fil. 1948;142:241–3.
2. Sorenson GD, Pribish DM, Valone FH, Memoli VA, Bzik DJ, Yao SL. Soluble normal and mutated DNA sequences from single-copy genes in human blood. Cancer Epidemiol Biomarkers Prev. 1994;3:67–71.
3. Nawroz H, Koch W, Anker P, Stroun M, Sidransky D. Microsatellite alterations in serum DNA of head and neck cancer patients. Nat Med. 1996;2:1035–7.
4. Stroun M, Lyautey J, Lederrey C, Olson-Sand A, Anker P. About the possible origin and mechanism of circulating DNA apoptosis and active DNA release. Clin Chim Acta. 2001;313:139–42.
5. Diaz LA Jr, Bardelli A. Liquid biopsies: genotyping circulating tumor DNA. J Clin Oncol. 2014;32:579–86.
6. Pantel K, Alix-Panabieres C. Circulating tumour cells in cancer patients: challenges and perspectives. Trends Mol Med. 2010;16:398–406.
7. Diehl F, Li M, Dressman D, He Y, Shen D, Szabo S, et al. Detection and quantification of mutations in the plasma of patients with colorectal tumors. Proc Natl Acad Sci U S A. 2005;102:16368–73.
8. Jahr S, Hentze H, Englisch S, Hardt D, Fackelmayer FO, Hesch RD, et al. DNA fragments in the blood plasma of cancer patients: quantitations and evidence for their origin from apoptotic and necrotic cells. Cancer Res. 2001;61:1659–65.
9. Sacher AG, Paweletz C, Dahlberg SE, Alden RS, O’Connell A, Feeney N, et al. Prospective validation of rapid plasma genotyping for the detection of EGFR and KRAS mutations in advanced lung cancer. JAMA Oncol. 2016;2:1014–22.
10. Diehl F, Schmidt K, Choti MA, Romans K, Goodman S, Li M, et al. Circulating mutant DNA to assess tumor dynamics. Nat Med. 2008;14:985–90.
11. Umetani N, Giuliano AE, Hiramatsu SH, Amersi F, Nakagawa T, Martino S, et al. Prediction of breast tumor progression by integrity of free circulating DNA in serum. J Clin Oncol. 2006;24:4270–6.
12. Li L, Hann HW, Wan S, Hann RS, Wang C, Lai Y, et al. Cell-free circulating mitochondrial DNA content and risk of hepatocellular carcinoma in patients with chronic HBV infection. Sci Rep. 2016;6:23992.
13. Lehmann-Werman R, Neiman D, Zemmour H, Moss J, Magenheim J, Vaknin-Dembinsky A, et al. Identification of tissue-specific cell death using methylation patterns of circulating DNA. Proc Natl Acad Sci U S A. 2016;113:E1826–34.
14. Schwarzenbach H, Hoon DS, Pantel K. Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer. 2011;11:426–37.
15. Xie M, Lu C, Wang J, McLellan MD, Johnson KJ, Wendl MC, et al. Age-related mutations associated with clonal hematopoietic expansion and malignancies. Nat Med. 2014;20:1472–8.
16. Genovese G, Kahler AK, Handsaker RE, Lindberg J, Rose SA, Bakhoum SF, et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N Engl J Med. 2014;371:2477–87.
17. Jaiswal S, Fontanillas P, Flannick J, Manning A, Grauman PV, Mar BG, et al. Age-related clonal hematopoiesis associated with adverse outcomes. N Engl J Med. 2014;371:2488–98.
18. Wan JC, Massie C, Garcia-Corbacho J, Mouliere F, Brenton JD, Caldas C, et al. Liquid biopsies come of age: towards implementation of circulating tumour DNA. Nat Rev Cancer. 2017;17:223–38.
19. Bettegowda C, Sausen M, Leary RJ, Kinde I, Wang Y, Agrawal N, et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med. 2014;6:224ra24.
20. Cisneros-Villanueva M, Hidalgo-Perez L, Rios-Romero M, Cedro-Tanda A, Ruiz-Villavicencio CA, Page K, et al. Cell-free DNA analysis in current cancer clinical trials: a review. Br J Cancer. 2022;126:391–400.
21. Pantel K, Alix-Panabieres C. Real-time liquid biopsy in cancer patients: fact or fiction? Cancer Res. 2013;73:6384–8.
22. Bronkhorst AJ, Ungerer V, Diehl F, Anker P, Dor Y, Fleischhacker M, et al. Towards systematic nomenclature for cell-free DNA. Hum Genet. 2021;140:565–78.
23. BioRender. Create professional science figures in minutes [Internet]. Toronto, ON: BioRender; 2022. [cited 2022 Oct 10]. Available from: https://biorender.com/
24. Keller L, Belloum Y, Wikman H, Pantel K. Clinical relevance of blood-based ctDNA analysis: mutation detection and beyond. Br J Cancer. 2021;124:345–58.
25. Gray ES, Witkowski T, Pereira M, Calapre L, Herron K, Irwin D, et al. Genomic analysis of circulating tumor DNA using a melanoma-specific UltraSEEK Oncogene Panel. J Mol Diagn. 2019;21:418–26.
26. Kohn L, Johansson M, Grankvist K, Nilsson J. Liquid biopsies in lung cancer-time to implement research technologies in routine care? Ann Transl Med. 2017;5:278.
27. Diehl F, Li M, He Y, Kinzler KW, Vogelstein B, Dressman D. BEAMing: single-molecule PCR on microparticles in water-in-oil emulsions. Nat Methods. 2006;3:551–9.
28. Sorber L, Zwaenepoel K, Deschoolmeester V, Van Schil PE, Van Meerbeeck J, Lardon F, et al. Circulating cell-free nucleic acids and platelets as a liquid biopsy in the provision of personalized therapy for lung cancer patients. Lung Cancer. 2017;107:100–7.
29. Santis G, Angell R, Nickless G, Quinn A, Herbert A, Cane P, et al. Screening for EGFR and KRAS mutations in endobronchial ultrasound derived transbronchial needle aspirates in non-small cell lung cancer using COLD-PCR. PLoS One. 2011;6:e25191.
30. Castellanos-Rizaldos E, Liu P, Milbury CA, Guha M, Brisci A, Cremonesi L, et al. Temperature-tolerant COLD-PCR reduces temperature stringency and enables robust mutation enrichment. Clin Chem. 2012;58:1130–8.
31. Narayan A, Carriero NJ, Gettinger SN, Kluytenaar J, Kozak KR, Yock TI, et al. Ultrasensitive measurement of hotspot mutations in tumor DNA in blood using error-suppressed multiplexed deep sequencing. Cancer Res. 2012;72:3492–8.
32. Newman AM, Lovejoy AF, Klass DM, Kurtz DM, Chabon JJ, Scherer F, et al. Integrated digital error suppression for improved detection of circulating tumor DNA. Nat Biotechnol. 2016;34:547–55.
33. Dressman D, Yan H, Traverso G, Kinzler KW, Vogelstein B. Transforming single DNA molecules into fluorescent magnetic particles for detection and enumeration of genetic variations. Proc Natl Acad Sci U S A. 2003;100:8817–22.
34. Dang DK, Park BH. Circulating tumor DNA: current challenges for clinical utility. J Clin Invest. 2022;132:e154941.
35. Chen M, Zhao H. Next-generation sequencing in liquid biopsy: cancer screening and early detection. Hum Genomics. 2019;13:34.
36. Mosko MJ, Nakorchevsky AA, Flores E, Metzler H, Ehrich M, van den Boom DJ, et al. Ultrasensitive detection of multiplexed somatic mutations using MALDI-TOF mass spectrometry. J Mol Diagn. 2016;18:23–31.
37. Arisi MF, Dotan E, Fernandez SV. Circulating tumor DNA in precision oncology and its applications in colorectal cancer. Int J Mol Sci. 2022;23:4441.
38. Baer C, Kern W, Koch S, Nadarajah N, Schindela S, Meggendorfer M, et al. Ultra-deep sequencing leads to earlier and more sensitive detection of the tyrosine kinase inhibitor resistance mutation T315I in chronic myeloid leukemia. Haematologica. 2016;101:830–8.
39. Soda N, Clack K, Shiddiky MJ. Recent advances in liquid biopsy technologies for cancer biomarker detection. Sens Diagn. 2022;1:343–75.
40. Underhill HR, Kitzman JO, Hellwig S, Welker NC, Daza R, Baker DN, et al. Fragment length of circulating tumor DNA. PLoS Genet. 2016;12:e1006162.
41. Chen A, Li J, Wang L, Huang Q, Zhu J, Wen S, et al. Comparison of paired cerebrospinal fluid and serum cell-free mitochondrial and nuclear DNA with copy number and fragment length. J Clin Lab Anal. 2020;34:e23238.
42. Cristiano S, Leal A, Phallen J, Fiksel J, Adleff V, Bruhm DC, et al. Genome-wide cell-free DNA fragmentation in patients with cancer. Nature. 2019;570:385–9.
43. Cai Z, Wang Z, Liu C, Shi D, Li D, Zheng M, et al. Detection of microsatellite instability from circulating tumor DNA by targeted deep sequencing. J Mol Diagn. 2020;22:860–70.
44. Arzimanoglou II, Gilbert F, Barber HR. Microsatellite instability in human solid tumors. Cancer. 1998;82:1808–20.
45. Cortes-Ciriano I, Lee S, Park WY, Kim TM, Park PJ. A molecular portrait of microsatellite instability across multiple cancers. Nat Commun. 2017;8:15180.
46. Boyiadzis MM, Kirkwood JM, Marshall JL, Pritchard CC, Azad NS, Gulley JL. Significance and implications of FDA approval of pembrolizumab for biomarker-defined disease. J Immunother Cancer. 2018;6:35.
47. Niu B, Ye K, Zhang Q, Lu C, Xie M, McLellan MD, et al. MSIsensor: microsatellite instability detection using paired tumor-normal sequence data. Bioinformatics. 2014;30:1015–6.
48. Salipante SJ, Scroggins SM, Hampel HL, Turner EH, Pritchard CC. Microsatellite instability detection by next generation sequencing. Clin Chem. 2014;60:1192–9.
49. Kautto EA, Bonneville R, Miya J, Yu L, Krook MA, Reeser JW, et al. Performance evaluation for rapid detection of pan-cancer microsatellite instability with MANTIS. Oncotarget. 2017;8:7452–63.
51. Luo H, Zhao Q, Wei W, Zheng L, Yi S, Li G, et al. Circulating tumor DNA methylation profiles enable early diagnosis, prognosis prediction, and screening for colorectal cancer. Sci Transl Med. 2020;12:eaax7533.
52. Ofman JJ, Hall MP, Aravanis AM. GRAIL and the quest for earlier multi-cancer detection [Internet]. Berlin: Springer Nature; 2018. [cited 2022 Oct 10]. Available from: https://www.nature.com/articles/d42473-020-00079-y
53. Koch A, Joosten SC, Feng Z, de Ruijter TC, Draht MX, Melotte V, et al. Analysis of DNA methylation in cancer: location revisited. Nat Rev Clin Oncol. 2018;15:459–66.
54. Daniunaite K, Jarmalaite S, Kriukiene E. Epigenomic technologies for deciphering circulating tumor DNA. Curr Opin Biotechnol. 2019;55:23–9.
55. Colella S, Shen L, Baggerly KA, Issa JP, Krahe R. Sensitive and quantitative universal pyrosequencing methylation analysis of CpG sites. Biotechniques. 2003;35:146–50.
56. Tost J, Dunker J, Gut IG. Analysis and quantification of multiple methylation variable positions in CpG islands by pyrosequencing. Biotechniques. 2003;35:152–6.
57. Wong IH, Lo YM, Zhang J, Liew CT, Ng MH, Wong N, et al. Detection of aberrant p16 methylation in the plasma and serum of liver cancer patients. Cancer Res. 1999;59:71–3.
58. Wojdacz TK, Dobrovic A, Hansen LL. Methylation-sensitive high-resolution melting. Nat Protoc. 2008;3:1903–8.
59. Xia Y, Tang W, Qian X, Li X, Cheng F, Wang K, et al. Efficacy and safety of camrelizumab plus apatinib during the perioperative period in resectable hepatocellular carcinoma: a single-arm, open label, phase II clinical trial. J Immunother Cancer. 2022;10:e004656.
60. Openshaw MR, Mohamed AA, Ottolini B, Fernandez-Garcia D, Richards CJ, Page K, et al. Longitudinal monitoring of circulating tumour DNA improves prognostication and relapse detection in gastroesophageal adenocarcinoma. Br J Cancer. 2020;123:1271–9.
61. Martinez-Saez O, Pascual T, Braso-Maristany F, Chic N, Gonzalez-Farre B, Sanfeliu E, et al. Circulating tumor DNA dynamics in advanced breast cancer treated with CDK4/6 inhibition and endocrine therapy. NPJ Breast Cancer. 2021;7:8.
62. Engel T, Ben-Horin S, Beer-Gabel M. Autonomic dysfunction correlates with clinical and inflammatory activity in patients with Crohn’s disease. Inflamm Bowel Dis. 2015;21:2320–6.
63. Wang X, Liu H, Zhao C, Li W, Xu H, Chen Y. The DEAD-box RNA helicase 51 controls non-small cell lung cancer proliferation by regulating cell cycle progression via multiple pathways. Sci Rep. 2016;6:26108.
64. Merker JD, Oxnard GR, Compton C, Diehn M, Hurley P, Lazar AJ, et al. Circulating tumor DNA analysis in patients with cancer: American Society of Clinical Oncology and College of American Pathologists Joint Review. Arch Pathol Lab Med. 2018;142:1242–53.
65. El Messaoudi S, Rolet F, Mouliere F, Thierry AR. Circulating cell free DNA: preanalytical considerations. Clin Chim Acta. 2013;424:222–30.
66. Norton SE, Lechner JM, Williams T, Fernando MR. A stabilizing reagent prevents cell-free DNA contamination by cellular DNA in plasma during blood sample storage and shipping as determined by digital PCR. Clin Biochem. 2013;46:1561–5.
67. Toro PV, Erlanger B, Beaver JA, Cochran RL, VanDenBerg DA, Yakim E, et al. Comparison of cell stabilizing blood collection tubes for circulating plasma tumor DNA. Clin Biochem. 2015;48:993–8.
68. Barra GB, Santa Rita TH, de Almeida Vasques J, Chianca CF, Nery LF, Santana Soares Costa S. EDTA-mediated inhibition of DNases protects circulating cell-free DNA from ex vivo degradation in blood samples. Clin Biochem. 2015;48:976–81.
69. Li D, Kusko R, Ning B, Tong W, Johann DJ Jr, Xu J. FDA-led consortium studies advance quality control of targeted next generation sequencing assays for precision oncology. Precis Cancer Med. 2021;4:32.
70. Gong B, Li D, Kusko R, Novoradovskaya N, Zhang Y, Wang S, et al. Cross-oncopanel study reveals high sensitivity and accuracy with overall analytical performance depending on genomic regions. Genome Biol. 2021;22:109.
71. Deveson IW, Gong B, Lai K, LoCoco JS, Richmond TA, Schageman J, et al. Evaluating the analytical validity of circulating tumor DNA sequencing assays for precision oncology. Nat Biotechnol. 2021;39:1115–28.
72. Zhang Y, Blomquist TM, Kusko R, Stetson D, Zhang Z, Yin L, et al. Deep oncopanel sequencing reveals within block position-dependent quality degradation in FFPE processed samples. Genome Biol. 2022;23:141.
73. Willey JC, Morrison TB, Austermiller B, Crawford EL, Craig DJ, Blomquist TM, et al. Advancing NGS quality control to enable measurement of actionable mutations in circulating tumor DNA. Cell Rep Methods. 2021;1:100106.
74. U.S. Food Drug Administration. Drug Approvals and database [Internet]. Silver Spring MD: U.S. Food and Drug Administration; 2020. [cited 2020 Oct 10]. Available from: https://www.fda.gov/drugs/resources-information-approved-drugs/fda-approves-liquid-biopsy-ngs-companion-diagnostic-test-multiple-cancers-and-biomarkers
75. Duffy MJ, Crown J. Use of circulating tumour DNA (ctDNA) for measurement of therapy predictive biomarkers in patients with cancer. J Pers Med. 2022;12:99.
76. Ettinger DS, Wood DE, Aisner DL, Akerley W, Bauman JR, Bharat A, et al. NCCN guidelines insights: non-small cell lung cancer, version 2.2021. J Natl Compr Canc Netw. 2021;19:254–66.
77. Corcoran RB. Liquid biopsy versus tumor biopsy for clinical-trial recruitment. Nat Med. 2020;26:1815–6.
78. Ghosh RK, Pandey T, Dey P. Liquid biopsy: a new avenue in pathology. Cytopathology. 2019;30:138–43.
79. Marrugo-Ramirez J, Mir M, Samitier J. Blood-based cancer biomarkers in liquid biopsy: a promising non-invasive alternative to tissue biopsy. Int J Mol Sci. 2018;19:2877.
80. Chae YK, Oh MS. Detection of minimal residual disease using ctDNA in lung cancer: current evidence and future directions. J Thorac Oncol. 2019;14:16–24.
81. Moding EJ, Nabet BY, Alizadeh AA, Diehn M. Detecting liquid remnants of solid tumors: circulating tumor DNA minimal residual disease. Cancer Discov. 2021;11:2968–86.
82. Pellini B, Chaudhuri AA. Circulating tumor DNA minimal residual disease detection of non-small-cell lung cancer treated with curative intent. J Clin Oncol. 2022;40:567–75.
83. Parikh AR, Van Seventer EE, Siravegna G, Hartwig AV, Jaimovich A, He Y, et al. Minimal residual disease detection using a plasma-only circulating tumor DNA assay in patients with colorectal cancer. Clin Cancer Res. 2021;27:5586–94.
85. Dresser R. Research ethics. Aligning regulations and ethics in human research. Science. 2012;337:527–8.
87. Sturgeon CM, Diamandis E. Laboratory medicine practice guidelines. Use of tumor markers in clinical practice: quality requirements. Washington, DC: National Academy of Clinical Biochemistry; 2009.
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||