AbstractPurpose
PIK3CA, encoding for subunit p110a of phosphatidylinositol 3 kinase, is frequently mutated in breast cancer. PIK3CA mutation was predictive for pathological complete response (pCR) in human epidermal growth factor 2 positive breast cancer. This study explores the association of PIK3CA mutation and pCR in triple-negative breast cancer (TNBC) treated with neoadjuvant chemotherapy.
Materials and MethodsA total of 92 patients with TNBC derived from a prospectively randomized phase II trial GeparSixto study (NCT01426880). Exon 9 and exon 20 of PIK3CA mutations were evaluated by using classical Sanger method with formalin-fixed paraffin-embedded tumor tissues.
ResultsSeven of 90 tumors (7.8%) were detectable with a PIK3CA H1047R mutation. Overall, PIK3CA H1047R mutation was significantly associated with a lower pCR rate (14.3% vs. 56.6%; odds ratio, 0.128; 95% confidence interval [CI], 0.015 to 1.108; p=0.047). In carboplatin-containing treatment patients, H1047R mutation trended to predict a lower pCR rate (20% vs. 62.5%; p=0.146). In a multivariable analysis, H1047R mutation trended to predict a lower pCR rate (hazard ratio, 0.1; 95% CI, 0.01 to 1; p=0.056).
IntroductionTriple-negative breast cancer (TNBC) which accounts for 10%-20% of all breast cancers, is characterized by lack of expression of estrogen receptor, progesterone receptor, and human epidermal growth factor 2 (HER2) [1]. Patients diagnosed with TNBC have a higher death rate within 5 years after diagnosis than other subtypes of breast cancer [2]. However, there is no specific agent against TNBC. Thus, identification and evaluation of new biomarkers and therapeutic agents is a high priority.
Phosphatidylinositol 3 kinase (PI3K)/mammalian target of rapamycin (mTOR) signaling pathway is one of the most important intracellular pathways, which is frequently activated in diverse cancers, also including breast cancer [3,4]. PIK3CA which encodes p110α, the predominant isoform of the catalytic subunit of class 1A phosphatidylinositol 3-kinase, is mutated in various human cancers [5]. More than 80% of PIK3CA mutations are found in one of three hot spots. Two of the hotspots, represented by the single amino acid substitutions E542K and E545K, are localized in the helical domain of the protein; the third one, represented by the H1047R substitution, resides in the kinase domain. These hotspot mutations in the helical or kinase domain of the PIK3CA gene induce oncogenic transformation through increasing the lipid kinase activity of p110a and gain function of PI3K [6,7].
PIK3CA mutations were reported in 25%-46% of primary breast cancer [8-11]. In mammary epithelial cells overexpression of PIK3CA hotspot mutant results in constitutive PI3K activation and induces multiple phenotypic alterations of breast cancer cells, including anchorage-independent proliferation in soft agar, growth factor-independent proliferation, and protection from anoikis. Moreover, these mutations also confer increased resistance to paclitaxel and induce abnormal mammary acinar morphogenesis in three-dimensional basement membrane cultures [6]. Recently, a meta-analysis by Loibl et al. [12] evaluated the association between PIK3CA mutation and response to anti-HER2 therapy in five studies with a total of 967 tumors. The results indicated that the presence of PIK3CA mutation was associated with a lower pCR rate in patients treated with target therapy against HER2, especially with double HER2 blockade (trastuzumab plus lapatinib) [12]. Furthermore, PIK3CA mutation was reported to be an independent factor for worse overall survival (OS) in patients with HER2-negative breast cancer [13]. However, the predictive role of PIK3CA mutation in TNBC still needs to be elucidated.
Pathological complete response (pCR) has predicted long-term outcome in several neoadjuvant studies and is therefore a potential surrogate marker for survival [14]. To better correlate pCR with disease-free survival and OS, a pooled analysis of several neoadjuvant trials has been published recently [15]. This large scale study showed pCR that was defined as no invasive residuals in breast tissue and lymph nodes (ypT0/is ypN0), as well as no invasive or non-invasive residual in breast or lymph nodes (ypT0 ypN0) was proposed as a surrogate endpoint for prediction of long-term clinical benefit in breast cancer. Furthermore, pCR defined as ypT0 (/is) ypN0 was strongly associated with long-term outcomes in patients with TNBC and in patients with HER2-positive, hormone receptor–negative tumors who received trastuzumab.
This study investigates the PIK3CA mutation and tries to find out the association between PIK3CA mutations and pCR rates in patients with TNBC treated with anthracycline-based neoadjuvant chemotherapy with or without carboplatin.
Materials and Methods1. Patient and treatmentIn present study, all patients are from a prospectively randomized phase II trial GeparSixto (NCT01426880) investigating the addition of carboplatin to neoadjuvant therapy for triple-negative and HER2-positive early breast cancer.
In GeparSixto study, women with previously untreated, unilateral or bilateral, centrally confirmed HER2-positive or triple-negative primary invasive breast cancer were enrolled after providing written informed consent. All patients were scheduled to receive paclitaxel 80 mg/m2 plus nonpegylated liposomal doxorubicin (Myocet, Elan Pharmaceuticals, Princeton, NJ): 20 mg/m2, both administered once per week for 18 weeks (PM). Patients with TNBC received bevacizumab 15 mg/kg intravenously every 3 weeks simultaneously with all cycles. Patients were randomly assigned to simultaneously receive carboplatin at a dose of area under the curve 2.0, which was reduced to the area under the curve 1.5 after accrual of 330 patients, once per week for 18 weeks (PMCb) or no additional treatment (PM only).
HER2 status (immunohistochemistry 1+ or in situ hybridization ratio < 2.2) and hormone receptor status (< 1%) were assessed as TNBCs in a central laboratory before random assignment. The current study selected 92 pretherapeutic tumor samples with TNBC from patients treated in the GeparSixto study for PIK3CA mutation evaluation. The characteristics of patients and lymphocyte infiltrating status were all collected from GeparSixto study. Based on a previous study [16], tumor with equal to or more than 60% of either intratumoral or stromal tumor-infiltrating lymphocytes was defined as lymphocyte-predominant breast cancer (LPBC).
2. PIK3CA mutation assessmentTumor materials were from formalin-fixed paraffin-embedded core biopsies which were taken before therapy. The tumor samples with a tumor content of equal to or more than 20% were used for PIK3CA mutation analysis.
The method which was used to detect PIK3CA mutations was described by Loibl et al. [17]. Magnetic beads-based DNA isolation from tissue sections was carried out by using the Siemens Versant system (Siemens Healthcare Diagnostics Inc., Tarrytown, NY). The following oligonucleotides were used as primers: exon 9 forward 5′-GGGAAAAATATGACAAAGAAAGC-3′, reverse 5′-GAGATCAGCCAAATTCAGTT-3′, exon 20 part 1 forward 5′-CATTTgCTCCAAACTGACCA-3′, reverse 5′-TGTGCATCATTCATTTGTTTCA-3′, exon 20 part 2 forward 5′-TTGATGACATTGCATACATTCG-3′, and reverse 5′-GGTCTTTGCCTGCTGAGAGT-3′. Polymerase chain reaction (PCR) was performed under standard conditions (94°C for 10 minutes, 94°C for 40 seconds, specific annealing temperature 56°C for exon 9 or 55°C for exon 20 for 1 minute, 72°C for 1 minute for 42 cycles; and 72°C for 10 minutes) by using 250 ng of template DNA in a 25 μL PCR volume. The PCR products were purified by using a Qiagen kit (Hilden, Germany), and DNA quality was assessed by using agarose gel electrophoresis and staining with ethidium bromide. The sequencing reaction of forward and reverse strands was performed on a 3100 Avant Genetic Analyzer (Applied Biosystems, Foster City, CA) by using 5 pM of each primer as specified previously and Big Dye Terminator Cycle Sequencing Mix v1.1 (Applied Biosystems). Sequencing alignment and quantification of mutated sequences were carried out by using BioEdit 7.1 software (Abbott Laboratories, Carlsbad, CA).
3. Statistical analysisAssociations between PIK3CA mutation, clinicopathological characteristics, and pCR were investigated with Fisher exact test or chi-square test for categorical variables. Odds ratios (ORs) and 95% confidence interval (CI) with two-sided p-values were used. In addition, a multivariable logistic regression model was constructed to adjust for known baseline characteristics such as age, tumor stage, nodal stage, histologic type, and therapy and to compare the predictive value of markers for pCR. All p-values are two-sided, with a p ≤ 0.05 considered to be statistically significant. Statistical analysis was performed using SPSS ver. 21.0 (IBM Corp., Armonk, NY).
4. Ethical statementAll patients provided written informed consent for study participation and biomaterial collection. The relevant authorities and ethics committees approved the studies. This project belongs to the exploratory endpoint of GeparSixto study which was a registered multi-center clinical trial (NCT01426-880) and was approved by all sub-center ethics committees. This study is reported according to the REMARK (Reporting Recommendations for Tumor Marker Prognostic Studies) criteria [18].
Results1. Baseline characteristicsNinety-two patients were all derived from the GeparSixto study that comprised patients with operable breast cancer (clinical stage T2–T4a-d tumors or T1c tumors with either clinical or histological stage N+ disease). Characteristics of patients which are derived from the clinical study database are shown in Table 1. Eighty-eight of 92 patients (95.7%) were diagnosed with ductal-invasive breast cancer that is the most common breast cancer subtype. LPBC accounts for 31.5% of all breast cancers. Tumors were commonly observed with negative lymph node involvement, high tumor grade. Of 92 patients, 46 (50%) were with PM treatment and 46 (50%) were with PMCb treatment. Baseline characteristics were well balanced between the two treatment arms.
2. Frequency of PIK3CA mutation
PIK3CA genotyping was performed in exon 9 and exon 20 separately for every sample. Of 92 tumor samples, two samples in exon 20 and exon 9 were undetectable due to bad quality of DNA or unavailable sequencing results. Three cases in exon 9 and eight cases in exon 20 were detectable with a mutation, which resulted in a mutation rate of 12.2% in PIK3CA (exon 9 and 20). In addition, one case with a silent mutation in exon 20 (c.3075 C>T) (Fig. 1C) and one case with a single nucleotide mutation in non-coding sequence after stop codon of PIK3CA (c.3236 T>C) (Fig. 1G) were observed. In this study, only the missense PIK3CA mutations in exon 9 or exon 20 were introduced into the following data analyses. Of 11 missense mutations, one case with E542K mutation (Fig. 1E), one case with E545K mutation (Fig. 1F) and one case with E547K mutation (Fig. 1D) were detected in exon 9; seven cases with H1047R mutation (Fig. 1B), and one case with E1037G mutation (Fig. 1A) were detected in exon 20. PIK3CA H1047R mutation was the most common mutation with a mutation rate of 7.8% (7 of 90). PIK3CA H1047R mutation did not correlate with other clinicopathological characteristics (Table 2).
3. Association of PIK3CA mutation and pCRIn the current study, four of 11 patients with PIK3CA-mutant tumors (36.4%) achieved pCR, compared with 43 of 79 patients with the wildtype tumors (54.4%). No statistical difference for pCR was observed (p=0.335) (Table 3). Overall, seven of 90 patients with PIK3CA mutation were detected with a H1047R mutation. And only one of seven patients with H1047R mutation tumor (14.3%) achieved a pCR compared with patients with H1047 wildtype (pCR rate, 56.6%). PIK3CA H1047R mutation was significantly associated with a lower pCR rate in complete cohort (OR, 0.128; 95% CI, 0.015 to 1.108; p=0.047) (Table 4).
A subgroup analysis of the two therapy arms was performed. In PMCb treatment arm, two of seven patients with PIK3CA mutation tumor (28.6%) achieved a pCR compared with 24 of 38 patients without PIK3CA mutation tumor (63.2%; p=0.114) (Table 3); one of five patients with PIK3CA H1047R mutant tumors (20%) achieved a pCR compared with 25 of 40 patients with PIK3CA H1047 wildtype tumors (62.5%) (p=0.146) (Table 4). There was a trending that both PIK3CA mutation and H1047R mutation were associated with a lower pCR rate in carboplatin additional treatment arm (Tables 3 and 4). Subgroup analysis was not performed in PM treatment arm due to the small mutation number (only four tumors with a mutation and two of these four tumors with a H1047R mutation in PM treatment arm) (Table 4).
In a multivariable logistic regression model, clinicopathological characteristics and PIK3CA H1047R mutation were showed in the multivariable logistic regression model for predicting pCR (Table 5). After adjustment for age, histologic type, tumor stage, nodal status, LPBC and chemotherapy, PIK3CA H1047R mutation was trending to predicting for a lower pCR rate (HR, 0.10; 95% CI, 0.01 to 1; p=0.056). LPBC independently predicted pCR (HR, 3.82; 95% CI, 1.24 to 11.76; p=0.020).
DiscussionCurrent study investigated the association of PIK3CA mutations and pCR in TNBCs after anthracycline-taxane–based neoadjuvant chemotherapy. The results showed that H1047R mutation was the most frequent mutation among the PIK3CA hotspot mutations and acquired a mutation rate of 7.8% in TNBCs,which is consistent with the data from COSMIC database. Moreover, this study for the first time showed that PIK3CA H1047R mutation was significantly associated with a lower pCR rate in early TNBC after anthracycline-taxane–based neoadjuvant chemotherapy. In addition, LPBC independently predicted a higher pCR in TNBCs treated with taxane-based neoadjuvant chemotherapy, which is consistent with the results from the complete cohort of GeparSixto study [19].
The mutations of PIK3CA were reported to induce oncogenic transformation with high efficiency through increasing the lipid kinase activity of p110a and gain function of PI3K [6-8,20]. Literature has reported that mutated PIK3CA is related to a poor prognosis in patients with breast cancer after treatment with anti-HER2 agents [21,22]. More recently, our research group reported that a significantly increased response to nab-paclitaxel was observed only in PIK3CA wildtype BC, including TNBC, but not in PIK3CA mutation BC [23]. Although PIK3CA mutation (including mutations in exon 9 and exon 20) was not significantly associated with a lower pCR rate in present study, H1047R mutation, as the most frequent mutation in PIK3CA, was significantly associated with a lower pCR rate (Table 4). As it is known, E542K, E545K, and H1047R mutations, are the three most frequently observed PI3K mutations, and all of these three hotspot mutations lead to an enzymatic activity, transform cells in vitro and enhance tumorigenicity in xenograft models. However, the tumorigenic abilities of these three hotspot mutations are a little different. The PIK3CA H1047R mutation was demonstrated to induce heterogeneous mammary carcinomas with a shorter latency and a higher frequency than E545K or E54-2K mutation, which is further confirmed due to more efficient activation of downstream AKT signaling [6,24]. In addition, the final results of GeparSixto study showed addition of carboplatin to neoadjuvant PM significantly improved pCR (53.2% with carboplatin vs 36.9% without carboplatin) in TNBC [25]. The present study showed PIK3CA H1047R was significantly associated a lower pCR, and further study in PMCb therapy arm showed that the patients with a PIK3CA H1047 wildtype tumor achieved a pCR rate of 62.5% compared with 20% with a PIK3CA H1047R mutation tumor (Table 4).
Therefore, PIK3CA H1047R mutation is probably a stronger driver mutation than other hotspot mutations of PIK3CA and it is likely to drive primary resistance to anthracyclinebased chemotherapy especially in platinum-containing regimen, which resulted in a significantly lower pCR in PIK3CA H1047R mutation tumor. In this study, we found that the presence of PIK3CA H1047R mutations could be used to stratify patients into distinct response to anthracycline-taxane–based neoadjuvant chemotherapy. Additionally, the study by Janku et al. [26] showed that H1047R mutant was an independent factor predicting response to PI3K/mTOR signaling pathway inhibitors. Mutation of PIK3CA gene may lead to a therapy resistance mechanism. Further studies should focus on the development of selective inhibitors against H1047R mutant PI3K in order to conquer this subtype of breast cancer.
Limitation to the small size, the information about subgroup could not be fully analyzed. For example, the comparisons of predictive role for response to neoadjuvant chemotherapy among PIK3CA hotspot mutations were not carried out in the present study. Further studies with a large sample number on validation of the predictive role of different PIK3CA hotspot mutations in TNBC are required. However, the current study for the first time provides new evidence for association of PIK3CA mutation and response to neoadjuvant chemotherapy in TNBC.
In summary, this study showed that TNBC with a PIK3CA H1047R mutation was less likely to achieve pCR after anthracycline-based neoadjuvant chemotherapy. Development of selective inhibitors against H1047R mutant PI3K is probably a new way to conquer this subtype of breast cancer.
AcknowledgmentsThis translational study was performed within the EU-funded FP7 RESPONSIFY project. Nr. 278659. We gratefully acknowledge the technical support of Ines Koch, Petra Wachs, and Britta Beyer.
Fig. 1.
PIK3CA mutation genotyping. Nucleotide sequence of PIK3CA mutations are shown. (A) 1624G>A (p. E542K) in one tumor. (B) 1633G>A (p.E545K) in one tumor. (C) c.1639G>A (p.E547K) in one tumor. (D) c.3110A>G (protein E1037G) in one tumor. (E) c.3140A>G (p.H1047R) in seven tumors. (F) c.3075 T>C (p. T1025T) in 1 tumor. (G) 3236T>C in one tumor. No tumor was detected with two mutations. 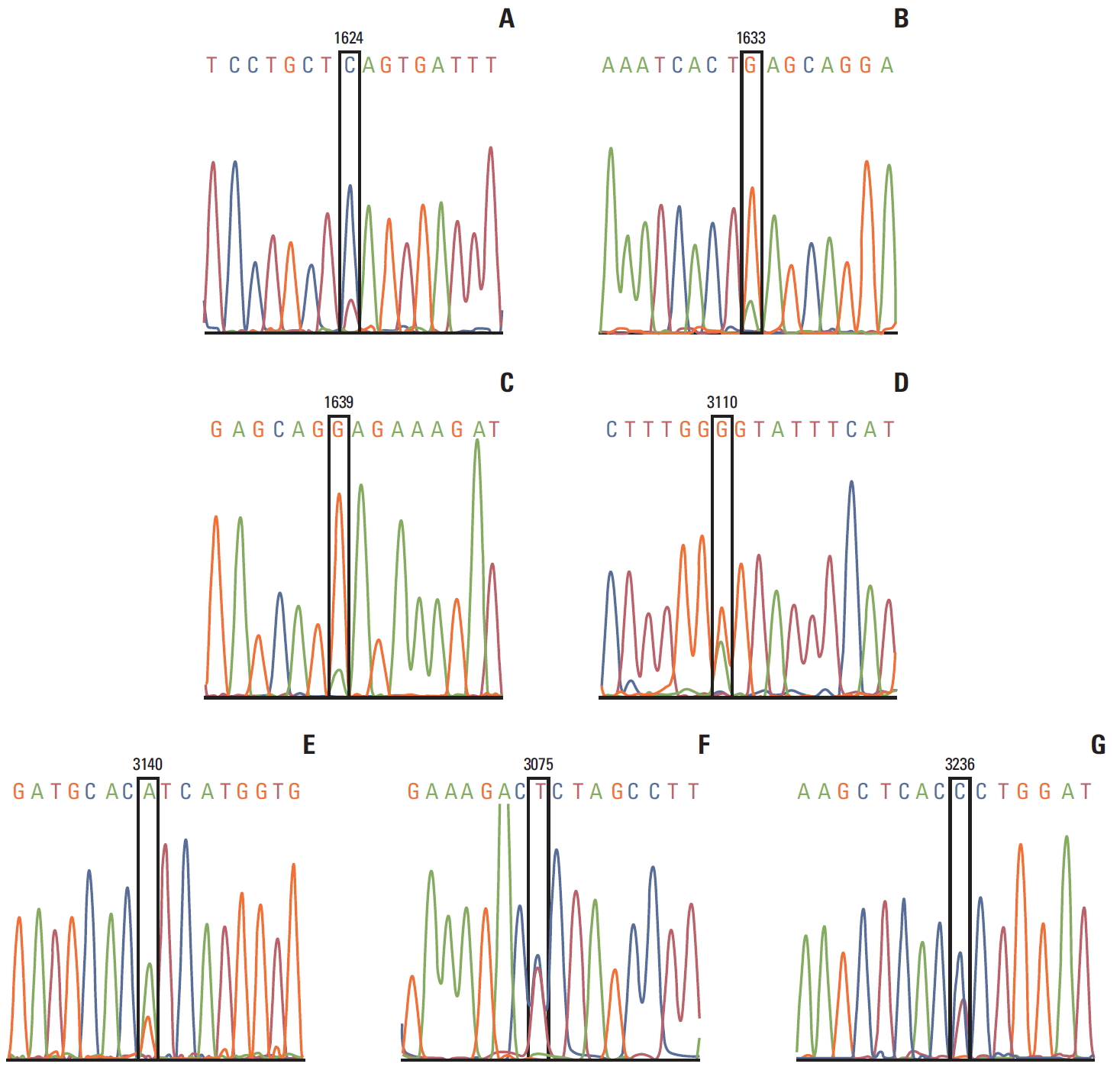
Table 1.Baseline characteristics of patients with early TNBC Table 2.Correlations of PIK3CA H1047R mutation and baseline characteristics in early TNBC (n=90) Table 3.pCR rates according to PIK3CA status overall and in two treatment arms separately
Table 4.pCR rates according to PIK3CA H1047R mutation overall and in two treatment arms separately
Table 5.Multivariable analysis of factors for predicting a pCR in early TNBC (n=92)
References1. Foulkes WD, Smith IE, Reis-Filho JS. Triple-negative breast cancer. N Engl J Med. 2010;363:1938–48.
2. Bauer KR, Brown M, Cress RD, Parise CA, Caggiano V. Descriptive analysis of estrogen receptor (ER)-negative, progesterone receptor (PR)-negative, and HER2-negative invasive breast cancer, the so-called triple-negative phenotype: a population-based study from the California cancer Registry. Cancer. 2007;109:1721–8.
3. Liu P, Cheng H, Roberts TM, Zhao JJ. Targeting the phosphoinositide 3-kinase pathway in cancer. Nat Rev Drug Discov. 2009;8:627–44.
4. Vinayak S, Carlson RW. mTOR inhibitors in the treatment of breast cancer. Oncology (Williston Park). 2013;27:38–44.
5. Samuels Y, Wang Z, Bardelli A, Silliman N, Ptak J, Szabo S, et al. High frequency of mutations of the PIK3CA gene in human cancers. Science. 2004;304:554.
6. Isakoff SJ, Engelman JA, Irie HY, Luo J, Brachmann SM, Pearline RV, et al. Breast cancer-associated PIK3CA mutations are oncogenic in mammary epithelial cells. Cancer Res. 2005;65:10992–1000.
7. Bader AG, Kang S, Vogt PK. Cancer-specific mutations in PIK3CA are oncogenic in vivo. Proc Natl Acad Sci U S A. 2006;103:1475–9.
8. Deng L, Zhu X, Sun Y, Wang J, Zhong X, Li J, et al. Prevalence and prognostic role of PIK3CA/AKT1 mutations in Chinese breast cancer patients. Cancer Res Treat. 2019;51:128–40.
9. Zardavas D, Marvelde L, Milne R, Joensuu H, Moynahan M, Hennessy B, et al. Tumor PIK3CA genotype and prognosis: a pooled analysis of 4,241 patients (pts) with early-stage breast cancer (BC). J Clin Oncol. 2015;33(15 Suppl):516.
10. Kalinsky K, Jacks LM, Heguy A, Patil S, Drobnjak M, Bhanot UK, et al. PIK3CA mutation associates with improved outcome in breast cancer. Clin Cancer Res. 2009;15:5049–59.
11. Campbell IG, Russell SE, Choong DY, Montgomery KG, Ciavarella ML, Hooi CS, et al. Mutation of the PIK3CA gene in ovarian and breast cancer. Cancer Res. 2004;64:7678–81.
12. Loibl S, Majewski I, Guarneri V, Nekljudova V, Holmes E, Bria E, et al. PIK3CA mutations are associated with reduced pathological complete response rates in primary HER2-positive breast cancer: pooled analysis of 967 patients from five prospective trials investigating lapatinib and trastuzumab. Ann Oncol. 2016;27:1519–25.
13. Li SY, Rong M, Grieu F, Iacopetta B. PIK3CA mutations in breast cancer are associated with poor outcome. Breast Cancer Res Treat. 2006;96:91–5.
14. Kaufmann M, Hortobagyi GN, Goldhirsch A, Scholl S, Makris A, Valagussa P, et al. Recommendations from an international expert panel on the use of neoadjuvant (primary) systemic treatment of operable breast cancer: an update. J Clin Oncol. 2006;24:1940–9.
15. Cortazar P, Zhang L, Untch M, Mehta K, Costantino JP, Wolmark N, et al. Pathological complete response and long-term clinical benefit in breast cancer: the CTNeoBC pooled analysis. Lancet. 2014;384:164–72.
16. Denkert C, Loibl S, Noske A, Roller M, Muller BM, Komor M, et al. Tumor-associated lymphocytes as an independent predictor of response to neoadjuvant chemotherapy in breast cancer. J Clin Oncol. 2010;28:105–13.
17. Loibl S, von Minckwitz G, Schneeweiss A, Paepke S, Lehmann A, Rezai M, et al. PIK3CA mutations are associated with lower rates of pathologic complete response to anti-human epidermal growth factor receptor 2 (her2) therapy in primary HER2-overexpressing breast cancer. J Clin Oncol. 2014;32:3212–20.
18. McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM, et al. REporting recommendations for tumor MARKer prognostic studies (REMARK). Breast Cancer Res Treat. 2006;100:229–35.
19. Denkert C, von Minckwitz G, Brase JC, Sinn BV, Gade S, Kronenwett R, et al. Tumor-infiltrating lymphocytes and response to neoadjuvant chemotherapy with or without carboplatin in human epidermal growth factor receptor 2-positive and triple-negative primary breast cancers. J Clin Oncol. 2015;33:983–91.
20. Zhao JJ, Liu Z, Wang L, Shin E, Loda MF, Roberts TM. The oncogenic properties of mutant p110alpha and p110beta phosphatidylinositol 3-kinases in human mammary epithelial cells. Proc Natl Acad Sci U S A. 2005;102:18443–8.
21. Baselga J, Cortes J, Im SA, Clark E, Ross G, Kiermaier A, et al. Biomarker analyses in CLEOPATRA: a phase III, placebo-controlled study of pertuzumab in human epidermal growth factor receptor 2-positive, first-line metastatic breast cancer. J Clin Oncol. 2014;32:3753–61.
22. Berns K, Horlings HM, Hennessy BT, Madiredjo M, Hijmans EM, Beelen K, et al. A functional genetic approach identifies the PI3K pathway as a major determinant of trastuzumab resistance in breast cancer. Cancer Cell. 2007;12:395–402.
23. Loibl S, Treue D, Budczies J, Weber K, Stenzinger A, Schmitt WD, et al. Mutational Diversity and therapy response in breast cancer: a sequencing analysis in the neoadjuvant GeparSepto trial. Clin Cancer Res. 2019;25:3986–95.
24. Meyer DS, Koren S, Leroy C, Brinkhaus H, Müller U, Klebba I, et al. Expression of PIK3CA mutant E545K in the mammary gland induces heterogeneous tumors but is less potent than mutant H1047R. Oncogenesis. 2013;2:e74
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||