AbstractPurposeThis preliminary study was conducted to evaluate the association between Oncotype DX (ODX) recurrence score and traditional prognostic factors. We also developed a nomogram to predict subgroups with low ODX recurrence scores (less than 25) and to avoid additional chemotherapy treatments for those patients.
Materials and MethodsClinicopathological and immunohistochemical variables were retrospectively retrieved and analyzed from a series of 485 T1-3N0-1miM0 hormone receptor-positive, human epidermal growth factor 2‒negative breast cancer patients with available ODX test results at Asan Medical Center from 2010 to 2016. One hundred twenty-seven patients (26%) had positive axillary lymph node micrometastases, and 408 (84%) had ODX recurrence scores of ≤25. Logistic regression was performed to build a nomogram for predicting a low-risk subgroup of the ODX assay.
ResultsMultivariate analysis revealed that estrogen receptor (ER) score, progesterone receptor (PR) score, histologic grade, lymphovascular invasion (LVI), and Ki-67 had a statistically significant association with the low-risk subgroup. With these variables, we developed a nomogram to predict the low-risk subgroup with ODX recurrence scores of ≤25. The area under the receiver operating characteristic curve was 0.90 (95% confidence interval [CI], 0.85 to 0.96). When applied to the validation group the nomogram was accurate with an area under the curve = 0.88 (95% CI, 0.83 to 0.95).
ConclusionThe low ODX recurrence score subgroup can be predicted by a nomogram incorporating five traditional prognostic factors: ER, PR, histologic grade, LVI, and Ki-67. Our nomogram, which predicts a low-risk ODX recurrence score, will be a useful tool to help select patients who may or may not need additional ODX testing.
IntroductionUntil now, a combination of clinical and histopathologic prognostic factors such as age [1], tumor size [2], lymph node metastasis [3], proliferation rate [4], and histologic grade [5] of breast cancer patients have been determined to use whether to treat patients with systemic chemotherapy. We also routinely evaluate predictors such as estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor 2 (HER2) to find patients suitable for a particular target therapy [6]. These are useful biomarkers to identify high-risk, triple-negative or HER2-positive cases. However, it is still difficult to accurately assess the individual risk and the need for accurate systemic chemotherapy in each case for low-risk patients among hormone receptor-positive patients. In the early 2000s, adjuvant chemotherapy was given to most early breast cancer patients, and such recommendations were made for hormone receptor positive patients, and with the development of multiple genetic or immunohistochemistry-based scores, more accurate clinical decisions have become possible. Oncotype DX (ODX) is a multigenic quantitative reverse transcription-polymerase chain reaction assay that predicts the efficacy of chemotherapy in patients with ER-positive breast cancer and assesses the risk of systemic recurrence. This has proven to be effective in predicting the effect and prognosis of chemotherapy in patients with breast cancer who are negative for ER-positive lymph nodes [7,8]. However, this test takes a long time to get the results and is expensive ($3,800). Because the test is complexed and the sample of the patient's tissue needs to be sent to the laboratory in order to obtain the results, Korea's National Health Insurance or any other private health insurance does not cover the ODX test. Therefore, this test is a costly burden for breast cancer patients, whether or not they are enrolled in private health insurance.
If other tests can accurately predict the recurrence score (RS) of all ER-positive patients, ODX testing will not be necessary Therefore, according to several studies, the information by ODX RS and the standard histopathological variables showed similar results [9,10]. However, there are many limitations to the results reported so far.
Our hypothesis was that risk assessments by ODX for ER-positive tumors, whether low or high risk, are predicted with high confidence. Therefore, the purpose of this preliminary study is to assess the association between traditional prognostic factors and ODX RSs and to develop a nomogram that can predict subgroups with low ODX RSs (less than 25) that can avoid additional chemotherapy.
Materials and Methods1. Patient selection and pathology variable selectionThe primary cohort of this study was selected through an evaluation of the initial record review of all T1-3N0-miM0 hormone receptor-positive, HER2-negative breast cancer patients with tumor tissue analyzed by the 21-gene assay between January 2010 and October 2016 at Asan Medical Center. A study data set of 485 cases with available ODX test results was used to build the prediction models. From January 2010 to December 2011, an independent external validation cohort of 1,166 consecutive patients was selected using the same criteria as the primary cohort. The clinical information of the patients was extracted from the medical record. Clinical information included patient age, tumor size, lymph node (LN) status, pathologic stage, histologic grade, nuclear grade, lymphovascular invasion (LVI), Ki-67, p53, and molecular subtypes according to ER, PR, and HER2 status based on immunohistochemistry or fluorescence in situ hybridization. Immunohistochemistry for ER, PR, HER2, and Ki-67 and in situ hybridization testing for HER2 were performed in the Asan Medical Center pathology laboratories. Nuclear staining for ER and PR was evaluated by the Allred scoring method (0-8). Membranous staining for HER2 was evaluated by the HercepTest (BenchMark XT autostainer using OptiView DAB Detection Kit, Ventana Medical Systems, Tucson, AZ) protocol. Immunohistochemical staining for Ki-67 (1:250, MIB-1, Dako, Glostrup, Denmark) was performed in a BenchMark XT autostainer (Ventana Medical Systems) using an i-View detection kit (Ventana Medical Systems).
2. Statistical analysisFor this study, we adopted an RS cutoff of 25 as the criterion for high risk, based on the RS threshold for chemotherapy administration that is used in ongoing phase III clinical trials such as TAILORx (Trial Assigning Individualized Options for Treatment) and RxPONDER (Rx for Positive Node, Endocrine Responsive Breast Cancer) after reanalysis of NSABP-B20. Chi-square test and Fisher exact test were used to compare the RS results between clinicopathological characteristics, when appropriate. Initial variable selection was carried out on the basis of univariate linear regression in development samples. Stepwise multiple logistic regression analysis was used to estimate a predictive model for ODX RS. Five factors such as nuclear grade (range, 1 to 3), Ki-67 labeling index (0 to 100), LVI (positive or negative), and Allred scores (range, 0 to 8) for ER and PR were found to contribute significantly to the multivariate logistic regression model. We conducted a robustness analysis to validate our model by employing the random sampling validation procedure. We randomly partitioned the data into two subsets where the sample size was approximately 340 at a ratio of 7:3, ran each random effect logistic regression using training set and test set, and employed receiver operating characteristic curve analysis and calculated the area under the curve (AUC). Using the Kaplan-Meier method to externally validate our nomogram system, we generated survival curves for breast cancer patients from 2010 to 2011. The significance of differences in survival was tested by using the log-rank test. All data analyses were performed using R statistical package ver. 3.2.0 (http://r-project.org). A significance level was set at 0.05 and all p-values reported were two-sided.
Results1. Patient characteristicsOf the 2,815 patients with pT1-3N0-1miM0, hormone receptor+/HER2− breast cancer included in this study, 485 patients (17%) with tumor tissue analyzed by the 21-gene assay were used to develop the nomogram to predict a subgroup of patients with low ODX RSs (≤ 25). Detailed clinical characteristics of patients included in the discovery cohort are shown in Table 1. The average age of subjects was 47.9 years (range, 27 to 77 years), and the average tumor size was 1.9 cm (range, 0.1 to 7.2 cm). Four hundred fifty-six cases (94%) were invasive ductal, 18 cases were invasive lobular (4%), and 11 cases (2%) exhibited other features. Among 11 cases, four cases were mucinous cancer, three cases were invasive micropapillary carcinoma, two cases were tubulolobular carcinoma, one case was invasive cribriform carcinoma, and one case was encapsulated papillary carcinoma. Lymph-node metastasis was absent in 358 cases (74%) and present as micrometastasis in 127 cases (26%). Patients with pathologic stages from IA to IIIA were included and 42% of them were at stage IA. LVI was absent in 346 cases and present in 139 cases. Ki-67 was < 20 in 289 cases and > 20 in 196 cases. S1 Fig. shows the distribution of RSs in this study group. The average of RSs was 18 and ranged from 0 to 72.
Table 1 also shows the characteristics of the study population included in this analysis, in comparison with the characteristics of patients with tumors associated with 0-25 RS (low-risk cohort) and those with scores of 25-100 (high-risk cohort). Among the 485 patients, 408 (84.1%) had a low RS of ≤ 25, and 77 (15.9%) had a high RS of > 25. Histologic grade, nuclear grade, presence of lymphovascular involvement, ER status, PR status, and Ki-67 were statistically related to the RS results, but no statistical correlation was found between the tumor size and presence of micrometastasis with RS results. On the other hand, there was no statistically significant difference between these two groups in histologic type. Invasive ductal carcinomas had RSs in all risk categories, whereas all invasive lobular carcinomas were assigned to the low-risk group. Mean RS increased with increasing tumor grade. All nuclear grade 1 tumors (n=12) were in the low-risk RS category. Nuclear grade 3 tumors were spread throughout the risk groups.
2. Development of a prediction modelOur goals were to evaluate the association between the ODX RS and routine clinical and pathologic measures, and to develop a nomogram that could predict a subgroup of patients with low ODX RSs (≤ 25), in whom addition of chemotherapy can be avoided. We randomized the data into two groups of randomly sampled sizes of about 340 at a ratio of 7:3 to develop nomograms. Detailed clinical characteristics of patients included in the training cohort and validation cohort are shown in Table 2. There were no significant differences in clinical characteristics between these two cohorts.
Univariate analysis revealed that histological and nuclear grade (Ki-67, ER, and PR) and LVI were associated with low-risk RS. Age, tumor size, lymph-node status, and HER2 status were not associated with RS. The Ki-67 was found to reach highest statistical significance in terms of association with low-risk RS (data not shown). When those five factors, excluding histologic grade, were included in multivariate analysis, nuclear grade, Ki-67, ER, PR, and LVI were all found to be independent predictors of low-risk RS in this population. Histological grade was a statistically significant variable in univariate analyses but was highly correlated with the nuclear grade, and thus excluded from multivariate analyses for statistical analysis stability. The odds ratio and β coefficient associated with each significant factor in the multivariate model are shown in Table 3: ER and PR status were positive effect to low-risk RS, and nuclear grade, Ki-67, and LVI were negative effect to low-risk RS. The significant variables of the explanatory model were used to develop a nomogram to predict low-risk RS of ODX (Fig. 1). Less than 0.1 and 0.9 or more were not displayed. The total scores for individual patients ranged from 30 to 265. The overall predictive accuracy of the nomogram as measured by the AUC was 0.90 (95% confidence interval [CI], 0.85 to 0.96) for the training dataset of 340 patients, and 0.88 (95% CI, 0.83 to 0.95) for the validation dataset of 145 patients (Fig. 2).
Table 4 shows the sensitivity, specificity, and positive and negative predictive values according to each cutoff value. A positive predictive value refers to the ability to predict a low-risk RS without ODX testing. When using a probability of 90 as the cutoff value, 242 of the 249 patients (97%) with such probabilities were predicted to be low-risk RS. The specificity rate was also determined to investigate the clinical utility of the nomogram. When conservatively defining a cutoff value, the general significance level is 0.05. A cutoff value of 5% was used, as it was important to evaluate if the nomogram could correctly classify the patients with a low predicted probability of low-risk RS. Among 263 patients predicted to have a probability of low-risk RS of ≥ 97, three patients had a high-risk of RS; this corresponds to a 5% specificity rate.
3. Validation of nomogram in an independent cohortTo validate the nomogram, we performed an independent external validation study using data from a cohort of 1,202 patients from 2010 to 2011. Median follow-up time of this group was 74 months. Table 5 shows the clinicopathologic characteristics of the external validation cohort. In general, distributions were similar, but the external validation group tended to be patients with less aggressive cancer. This is because the group that performed ODX included many patients with more aggressive breast cancer, for whom oncologists had to make a treatment decision. Patients with lower-stage and lower-grade cancers were more likely to be included in the external validation cohort than in the discovery cohort.
Patients in the external validation cohort were classified into low-risk and high-risk groups according to the nomogram value of 97. This cutoff of 97 is a number with a specificity of 95, as described above. In external validation cohort, 673 patients (56.0%) were in the high-risk group with < 97 points and 529 (44.0%) in the low-risk group with ≥ 97 points. We assessed the impact of the nomogram on the prediction of recurrence and distant metastasis in the validation cohort. Kaplan-Meier analysis demonstrated that the low-risk group, according to the nomogram, had a significantly higher probability of recurrence (p=0.001) and distant metastasis (p=0.001) than did the high-risk group (Fig. 3). The 5-year distant metastasis-free rates for patients with nomogram low-risk and high-risk were 98.8% and 97.9%, respectively. This result demonstrated that the nomogram could clearly differentiate patients at high and low risk of distant metastasis. In the analysis of death, there was no statistical difference between the two groups. This is thought to be due to the fact that there were few death events. The 5-year overall survival rate for all patients was 99.1% (Fig. 3). However, since this external validation group included chemotherapy-treated patients, we performed multivariate analysis. In the multivariate analysis of disease-free survival and distant metastasis-free survival in patients with breast cancer, the hazard ratio in low-risk patients with 97 or higher based on score 97 was 0.361 (p=0.033) and 0.137 (p=0.024), respectively. The score 97 was a prognostic factor for disease-free survival and distant metastasis-free survival in externally validated patients (Table 6). Thus, we confirmed that the nomogram for predicting the ODX RS low risk can be distinguished from the risk of recurrence even in the external cohort.
4. User-friendly implementation of the predictive modelWe organized our nomogram model in a simple and intuitive online application, mobile application, and automatic calculator using Microsoft Excel worksheets (Fig. 4). The interface allows the user to input values for ER, PR, Ki-67, grade, and LVI. The standard output includes an estimate of the probability that the RS exceeds the low-risk threshold of 25. As such, our algorithm seems to identify a group of patients for whom ODX ordering is more likely to contribute information that is not already available using routine measures.
DiscussionThe ODX testing is the most widely used breast cancer genomic assay in the world. This test is supported by the American Society of Clinical Oncology and the National Comprehensive Cancer Network in treatment of node negative, ER-positive breast cancer patients [11,12], and is accepted as a useful test [7,8,13,14]. Although the ODX test has been widely implemented in the United States, it has become a heavy burden for many patients because of the high cost. Because ODX testing is so expensive, this test is currently only available in about one-third of breast cancer patients in the United States and about 20% in Europe [15,16]. Also, in Korea, this test is conducted only in foreign countries, so neither the National Health Insurance nor the private health insurance covers the ODX test. Therefore, the cost of ODX test is a burden in Korea regardless of whether breast cancer patients have private insurance or not. Several recent studies have reported that stratification of risk is possible more cost-effectively for certain patients by predicting ODX test results using histopathological variables from routine pathology test [17]. Clark et al. [18] and Zbytek et al. [19] reported that ODX RSs are highly dependent on parameters already used in routine pathological examinations. In addition, studies by Cuzick et al. [20] provide evidence that standard clinical and pathological parameters are superior to ODX analysis in predicting patients’ prognoses [20]. This study was performed to evaluate the ODX RS, which was based on routine standard histologic and immunohistochemical data. However, the best way to link this with clinical practice has not yet been proven. There is no test to accurately predict ODX RS, too. Nevertheless, some tools can help you decide whether or not you should perform an ODX test. We found that using this tool in combination with standard histopathological and immunohistochemical parameters was useful as an alternative to costly ODX testing for patients who were able to predict high- or low-risk through ODX testing.
There are five variables used in our model: ER, PR, Ki-67, grade, and LVI. These five clinicopathologic variables can be collected through any pathologic examination and are clinically established variables in predicting prognosis. In the present study, there were consistent significant differences for ER, PR, grade, LVI and Ki-67 between the ODX risk stratification categories, but not for HER2, presence of micrometastases, and tumor size. This suggests that a negative or equivocal HER2, tumor size, and presence of micrometastases may have less importance in risk stratification with ODX testing than ER, PR, grade, LVI or Ki-67. The HER2 value obtained from the ODX test performed in this study was the same as that reported in previous studies [21]. In the University of Tennessee Medical Center that developed a nomogram based on the National Cancer Database (27,719 ODX tested ER+/HER2–/LN- breast cancer patients) study, the tumor grade, PR, and LVI were found to have the same factor. In addition, this study also found that age, tumor size, and histological subtype as significant factors, which were not observed in the results of our study [22]. In the present study, age was not a significant factor; however, according to the results of a recently published TAILORx study, the chemotherapy benefit for invasive disease-free survival varied when the RS was combined with age (p=0.004), with some chemotherapy benefits found in women aged ≤ 50 years with a RS of 16-25. Therefore, additional research on younger patients is necessary [23].
The pathologic factor most closely related to ODX RS in the present study was Ki67 proliferation index. Williams et al. [24] reported that the expression of Ki-67 by immunohistochemistry was significantly correlated with RS, and Sahebjam et al. [25] also reported a 90% chance of having a high or intermediate ODX recurrence rate for tumors with a Ki-67 expression ≥ 25%. PR was also a major predictor of a high-risk or a low-risk ODX RS in our study Tang et al. [26] showed a correlation between the lack of PR expression and aggressive morphologic features, and Auerbach et al. [27] and Chaudhary et al [28]. confirmed recent observations that the ODX score was higher when PR was negative Clark et al. [18], who revealed the inverse relationship between PR expression level and ODX RS, regardless of tumor grade, also mentioned the importance of PR semiquantitative score. We found that tumor grade carried the high predictive value for a low-risk ODX score, same as previous study reported by Gage et al. [29]. It is not surprising that this grade, which has long been regarded by the Nottingham Prognostic Index as an important predictor of breast cancer prognosis, has been identified as a key predictor of the ODX score [17], which also was recognized at the St. Gallen Consensus Conference in 2011 [30]. LVI was also a major predictor of low-risk ODX recurrence in our study, but LVI in our model had little effect on the outcome of the ODX test. Our model used 25 as a cutoff of RS. Because the range of RSs used in trials such as TAILORx and RxPONDER currently in progress is different from the existing definition, patients with a RS of 25 or higher were classified as receiving chemotherapy, and the patients from 11 to 25 were randomly grouped, regarding the role of adjuvant chemotherapy. These studies identified the optimal cutoff values between patients in the intermediate (RS, 11-25) or low to intermediate (RS ≤ 25) risk groups with respect to the benefits of adjuvant chemotherapy [31].
The present study has several strengths. The nomogram produced by the study is user-friendly and available free of any cost to the user. The information needed for the nomogram is already generated by many pathology laboratories during the initial assessment of primary breast cancer. Pathologists and oncologists can easily calculate the expected RS and compare the results with the actual RS, by our Microsoft Excel worksheets, mobile applications, and online versions. If the expected RS is evident in the high-risk or low-risk group, the oncologist should not expect the outcome to differ significantly from the ODX. In these cases, clinicians can avoid using ODX. To develop a nomogram with a high and acceptable C index (0.85-0.89), 485 patients in the original cohort and 1,202 patients in the external validation cohort were used, respectively. Our external validation cohort has 95% specificity cutoff value, because whether patients with low-risk RS values can accurately be classified as low-risk patients remains to be assessed. Among the 263 patients with low-risk RS scores of ≥ 97, only 3 were high risk, which was consistent with 5% specificity. This is a valuable, large-scale study showing that clinicopathologic variables can be used for prediction of low-risk ODX RS using our nomogram models.
There are some limitations in this study. First, the population in this study was biased by the clinical selection of cases, because of this, the proportion of cases with low to intermediate RSs could be higher However, the distribution of scores observed in our study was similar that observed in commercial laboratories. This results in this study show that the distribution of risk groups in this study reflects clinical practice and supports the possibility of generalizing the findings. Therefore, the clinical case selection bias presented in this study may be regarded as a strength of this study because it reflects the usefulness of the actual application and testing. Second, the follow-up time for patients in our study (the 1,202 external validation cases were selected from 2010 to 2011) is short. Our early survival results need to be confirmed by longer follow up. By following clinical outcomes, we hope to verify the utility of our findings, and reinforce the importance of our nomogram. Moreover, Ki-67 measurement varied depending on other methods used. Therefore, the difference observed when using this nomogram can influence the decision.
Our nomogram, which predicts a low-risk ODX RS, will be a useful tool to help select patients who may or may not need additional ODX testing. It may also be a useful tool to replace ODX testing for patients who cannot afford the test, or for whom ODX testing is not available.
Electronic Supplementary MaterialSupplementary materials are available at Cancer Research and Treatment website (http://www.e-crt.org).
AcknowledgmentsThis study was supported by a grant (2015-668, 2016-668) from Asan Institute for Life Sciences, Asan Medical Center, Seoul, Korea.
Fig. 1.Nomogram to predict low-risk recurrence score of Oncotype Dx. LVI, lymphovascular invasion; ER, estrogen receptor; PR, progesterone receptor. 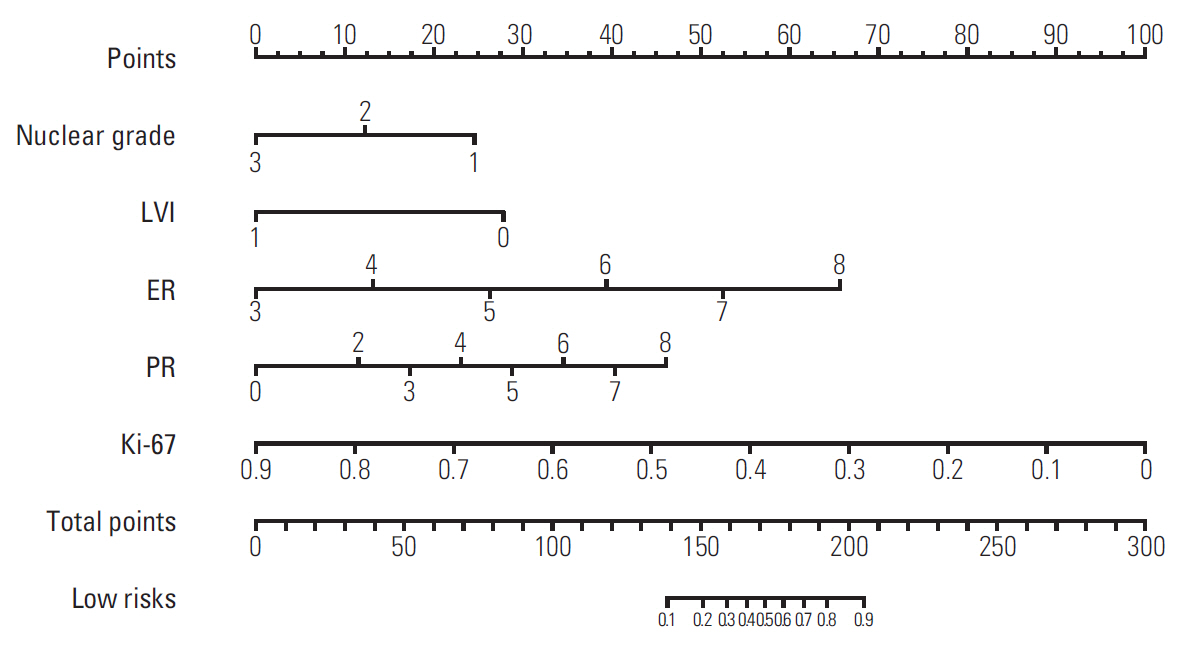
Fig. 2.Receiver operating characteristic curve (ROC) of nomogram. (A) Training dataset of 340 patients. (B) Validation dataset of 145 patients. 
Fig. 3.Kaplan-Meier analysis of external validation group according to cutoff of 97. (A) Disease-free survival. (B) Distant metastasis-free survival. (C) Overall survival. 
Fig. 4.Nomogram model. (A) Online application. (B) Mobile application. (C) Automatic calculator using Microsoft Excel worksheets. 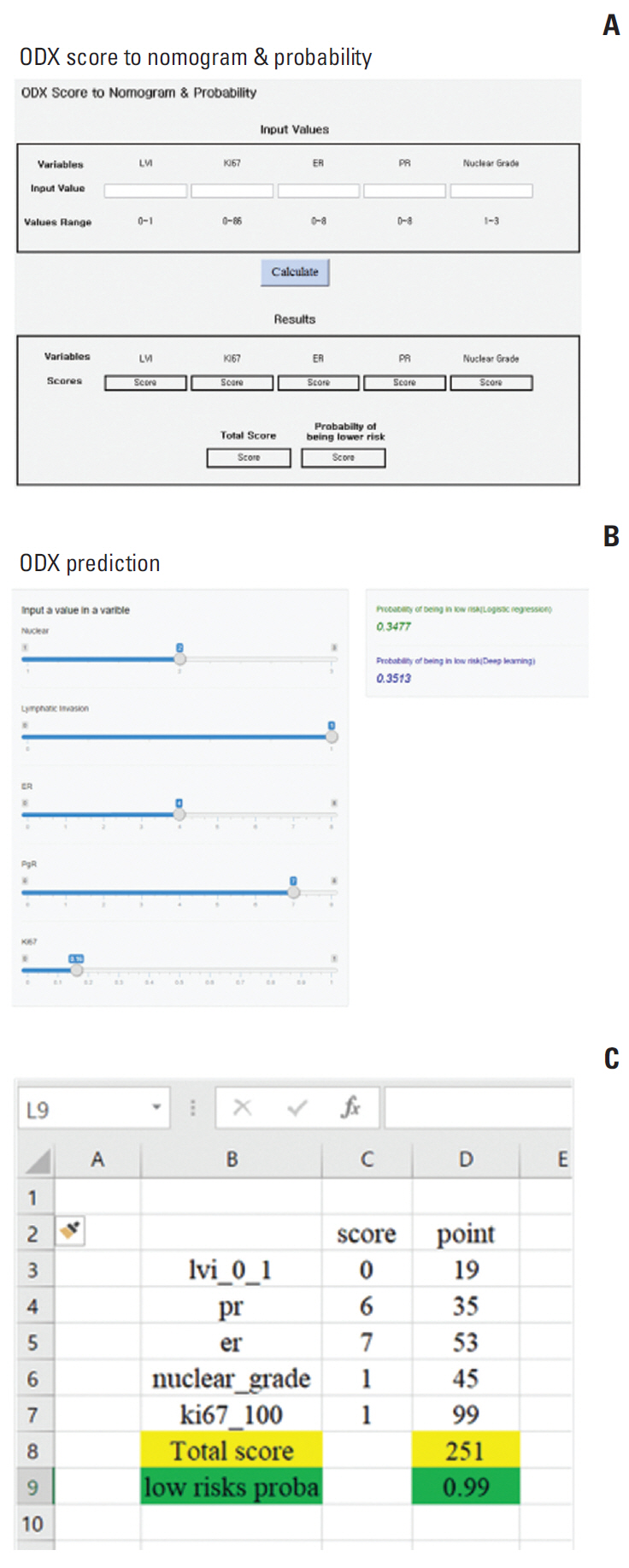
Table 1.Baseline characteristics and association between dichotomized Oncotype Dx score and clinicopathologic variables
Table 2.Clinicopathologic characteristics of the training group and internal validation group Table 3.Multivariate logistic regression model Table 4.Sensitivity, specificity, positive predictive and negative predictive values according to various cutoff values Table 5.Clinicopathologic characteristics of the external validation group Table 6.Multivariate analysis for DFS and DMFS References1. Rosenberg J, Chia YL, Plevritis S. The effect of age, race, tumor size, tumor grade, and disease stage on invasive ductal breast cancer survival in the U.S. SEER database. Breast Cancer Res Treat. 2005;89:47–54.
2. Koscielny S, Tubiana M, Le MG, Valleron AJ, Mouriesse H, Contesso G, et al. Breast cancer: relationship between the size of the primary tumour and the probability of metastatic dissemination. Br J Cancer. 1984;49:709–15.
3. Fisher B, Bauer M, Wickerham DL, Redmond CK, Fisher ER, Cruz AB, et al. Relation of number of positive axillary nodes to the prognosis of patients with primary breast cancer. An NSABP update. Cancer. 1983;52:1551–7.
4. Baak JP, Gudlaugsson E, Skaland I, Guo LH, Klos J, Lende TH, et al. Proliferation is the strongest prognosticator in node-negative breast cancer: significance, error sources, alternatives and comparison with molecular prognostic markers. Breast Cancer Res Treat. 2009;115:241–54.
5. Elston CW, Ellis IO. Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology. 2002;41:154–61.
6. Colditz GA, Rosner BA, Chen WY, Holmes MD, Hankinson SE. Risk factors for breast cancer according to estrogen and progesterone receptor status. J Natl Cancer Inst. 2004;96:218–28.
7. Joh JE, Esposito NN, Kiluk JV, Laronga C, Lee MC, Loftus L, et al. The effect of Oncotype DX recurrence score on treatment recommendations for patients with estrogen receptor-positive early stage breast cancer and correlation with estimation of recurrence risk by breast cancer specialists. Oncologist. 2011;16:1520–6.
8. Rouzier R, Pronzato P, Chereau E, Carlson J, Hunt B, Valentine WJ. Multigene assays and molecular markers in breast cancer: systematic review of health economic analyses. Breast Cancer Res Treat. 2013;139:621–37.
9. Klein ME, Dabbs DJ, Shuai Y, Brufsky AM, Jankowitz R, Puhalla SL, et al. Prediction of the Oncotype DX recurrence score: use of pathology-generated equations derived by linear regression analysis. Mod Pathol. 2013;26:658–64.
10. Kraus JA, Dabbs DJ, Beriwal S, Bhargava R. Semi-quantitative immunohistochemical assay versus oncotype DX((R)) qRT-PCR assay for estrogen and progesterone receptors: an independent quality assurance study. Mod Pathol. 2012;25:869–76.
11. Fitzgibbons PL, Dillon DA, Alsabeh R, Berman MA, Hayes DF, Hicks DG, et al. Template for reporting results of biomarker testing of specimens from patients with carcinoma of the breast. Arch Pathol Lab Med. 2014;138:595–601.
12. Harris L, Fritsche H, Mennel R, Norton L, Ravdin P, Taube S, et al. American Society of Clinical Oncology 2007 update of recommendations for the use of tumor markers in breast cancer. J Clin Oncol. 2007;25:5287–312.
13. Vataire AL, Laas E, Aballea S, Gligorov J, Rouzier R, Chereau E. Cost-effectiveness of a chemotherapy predictive test. Bull Cancer. 2012;99:907–14.
14. Blohmer JU, Rezai M, Kummel S, Kuhn T, Warm M, Friedrichs K, et al. Using the 21-gene assay to guide adjuvant chemotherapy decision-making in early-stage breast cancer: a cost-effectiveness evaluation in the German setting. J Med Econ. 2013;16:30–40.
15. Orucevic A, Heidel RE, Bell JL. Utilization and impact of 21-gene recurrence score assay for breast cancer in clinical practice across the United States: lessons learned from the 2010 to 2012 National Cancer Data Base analysis. Breast Cancer Res Treat. 2016;157:427–35.
16. Albanell J, Svedman C, Gligorov J, Holt SD, Bertelli G, Blohmer JU, et al. Pooled analysis of prospective European studies assessing the impact of using the 21-gene Recurrence Score assay on clinical decision making in women with oestrogen receptor-positive, human epidermal growth factor receptor 2-negative early-stage breast cancer. Eur J Cancer. 2016;66:104–13.
17. Carlson JJ, Roth JA. The impact of the Oncotype Dx breast cancer assay in clinical practice: a systematic review and meta-analysis. Breast Cancer Res Treat. 2013;141:13–22.
18. Clark BZ, Dabbs DJ, Cooper KL, Bhargava R. Impact of progesterone receptor semiquantitative immunohistochemical result on Oncotype DX recurrence score: a quality assurance study of 1074 cases. Appl Immunohistochem Mol Morphol. 2013;21:287–91.
19. Zbytek B, Cohen C, Wang J, Page A, Williams DJ, Adams AL. Nottingham-defined mitotic score: comparison with visual and image cytometric phosphohistone H3 labeling indices and correlation with Oncotype DX recurrence score. Appl Immunohistochem Mol Morphol. 2013;21:48–53.
20. Cuzick J, Dowsett M, Pineda S, Wale C, Salter J, Quinn E, et al. Prognostic value of a combined estrogen receptor, progesterone receptor, Ki-67, and human epidermal growth factor receptor 2 immunohistochemical score and comparison with the Genomic Health recurrence score in early breast cancer. J Clin Oncol. 2011;29:4273–8.
21. Chen LL, Nolan ME, Silverstein MJ, Mihm MC Jr, Sober AJ, Tanabe KK, et al. The impact of primary tumor size, lymph node status, and other prognostic factors on the risk of cancer death. Cancer. 2009;115:5071–83.
22. Orucevic A, Bell JL, McNabb AP, Heidel RE. Oncotype DX breast cancer recurrence score can be predicted with a novel nomogram using clinicopathologic data. Breast Cancer Res Treat. 2017;163:51–61.
23. Sparano JA, Gray RJ, Makower DF, Pritchard KI, Albain KS, Hayes DF, et al. Adjuvant chemotherapy guided by a 21-gene expression assay in breast cancer. N Engl J Med. 2018;379:111–21.
24. Williams DJ, Cohen C, Darrow M, Page AJ, Chastain B, Adams AL. Proliferation (Ki-67 and phosphohistone H3) and Oncotype DX recurrence score in estrogen receptor-positive breast cancer. Appl Immunohistochem Mol Morphol. 2011;19:431–6.
25. Sahebjam S, Aloyz R, Pilavdzic D, Brisson ML, Ferrario C, Bouganim N, et al. Ki 67 is a major, but not the sole determinant of Oncotype Dx recurrence score. Br J Cancer. 2011;105:1342–5.
26. Tang P, Wang J, Hicks DG, Wang X, Schiffhauer L, McMahon L, et al. A lower Allred score for progesterone receptor is strongly associated with a higher recurrence score of 21-gene assay in breast cancer. Cancer Invest. 2010;28:978–82.
27. Auerbach J, Kim M, Fineberg S. Can features evaluated in the routine pathologic assessment of lymph node-negative estrogen receptor-positive stage I or II invasive breast cancer be used to predict the Oncotype DX recurrence score? Arch Pathol Lab Med. 2010;134:1697–701.
28. Chaudhary LN, Jawa Z, Szabo A, Visotcky A, Chitambar CR. Relevance of progesterone receptor immunohistochemical staining to Oncotype DX recurrence score. Hematol Oncol Stem Cell Ther. 2016;9:48–54.
29. Gage MM, Rosman M, Mylander WC, Giblin E, Kim HS, Cope L, et al. A validated model for identifying patients unlikely to benefit from the 21-gene recurrence score assay. Clin Breast Cancer. 2015;15:467–72.
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||