AbstractPurposeMutation-specific antibodies have recently been developed for identification of epidermal growth factor receptor (EGFR) mutations by immunohistochemistry (IHC). This study was designed to investigate whether the type of specimen (biopsy vs. resection) would make a difference in determining mutation status by IHC, and to evaluate whether biopsies are suitable for detection of mutant EGFR protein.
Materials and MethodsIHC was performed using mutation-specific antibodies for E746-A750 deletion (DEL) and L858R point mutation (L858R) in biopsies and tissue microarrays of resected tumors from 154 patients with pulmonary adenocarcinoma. Results were then compared with DNA sequencing data.
ResultsMolecular-based assays detected EGFR mutations in 62 patients (40.3%), including 14 (9.1%) with DEL, and 31 (20.1%) with L858R. IHC with two mutation-specific antibodies showed a homogeneous staining pattern, and correctly identified EGFR mutation status in 89% (137/154). Overall (biopsy/resection) sensitivity, specificity, positive predictive value, and negative predictive value were 75.6% (78.3%/72.7%), 94.5% (90.9%/96.3%), 85% (78.3%/88.9%), and 90.4% (90.9%/89.7%), respectively.
ConclusionOur data showed that IHC using EGFR mutation–specific antibodies is useful for detection of EGFR mutations with high specificity and good sensitivity not only for resection specimens but also for biopsy materials. Therefore, IHC using EGFR mutation–specific antibodies may preclude a second biopsy procedure to obtain additional tissues for identification of EGFR mutations by molecular assays in biopsies from advanced cancer, particularly when tumor cells in the samples are limited.
IntroductionMutations in the tyrosine kinase domain of the epidermal growth factor receptor (EGFR) gene have been identified in non-small cell lung carcinoma (NSCLC), and they were correlated with clinical response to EGFR–tyrosine kinase inhibitor (TKI) treatment. Patients with EGFR mutations have a higher response rate to EGFR-TKIs (60%-80%) than those with EGFR–wild type or unknown mutation status (10%-20%) [1]. In addition, EGFR-TKI has been shown to be superior to carboplatin-paclitaxel as initial treatment [2]. DNA-based molecular methods have been used for selection of patients who would benefit from TKI therapy; however, they are tedious, expensive, and not routine in clinical laboratories. Yu et al. [3] recently generated mutation-specific rabbit monoclonal antibodies against the two most common NSCLC-associated EGFR mutations (E746-A750 deletion [DEL] in exon 19 and L858R point mutation [L858R] in exon 21) for detection of mutant EGFR protein by immunohistochemistry (IHC). IHC using these two mutation-specific antibodies has been suggested as a reliable screening method for EGFR-TKI therapy [4-8]. However, the immunostaining patterns reported in previous publications were contradictory; some heterogeneous, while others were described as homogeneous [9-11]. Most investigations used resected tumor specimens. Biopsy samples, however, may be the only tumor materials available for confirming EGFR mutation status, particularly in patients whose lung cancer is at an advanced stage and unresectable. In addition, tumor cells in the samples are often low in quantity or of insufficient quality for molecular assays, leading to a second biopsy procedure for obtaining additional tissues. It is therefore important to determine whether biopsy samples are suitable for detection of mutant EGFR protein.
There has been significant interest in total EGFR (tEGFR) protein expression, in particular, before the development of mutation-specific antibodies, however, its clinical significance is controversial. Some studies reported a better outcome after EGFR-TKI treatment for tumors overexpressing the tEGFR protein, whereas other studies did not [12-14]. To date, the literature contains limited data regarding the association between tEGFR and mutant EGFR proteins.
In the current study, immunohistochemical analysis of both biopsies and resected tumor tissues from lung adenocarcinoma with known EGFR mutation status by direct DNA sequencing was performed using mutation-specific antibodies against EGFR with E746-A750 deletion in exon 19 and L858R mutation in exon 21. Expression of tEGFR was also investigated in order to determine possible correlations with mutant EGFR proteins detected by mutation-specific antibodies.
Materials and Methods1. Patient characteristicsA total of 154 patients who underwent biopsy (n=78) or surgical resection (n=76) for pulmonary adenocarcinoma at the Catholic University St. Vincent’s Hospital from 2005 to 2009 were enrolled in this study. All cases had been previously screened for EGFR mutations by direct DNA sequencing. Paraffin-embedded tissues were procured from biopsied or resected specimens in a blinded fashion to the clinical information and EGFR mutation status, and IHC was performed using EGFR mutation–specific antibodies. The study protocol was approved by the Institutional Review Board (IRB) of St. Vincent’s Hospital at The Catholic University of Korea (IRB No. VC13TISI0055). Informed consent was waived by the IRB. Clinical information was obtained through a computerized retrospective database of the tumor registry. Clinicopathologic characteristics are summarized in Table 1.
2. Direct DNA sequencingThree 10-μm thick sections were cut from the paraffin blocks. Tissue corresponding to the precisely identified tumor areas on the hematoxylin and eosin (H&E) slides was scraped for subsequent DNA extraction using a QIAmp kit (Qiagen, Valencia, CA). Exons 18 to 21 of the TK domain in the EGFR gene were amplified by polymerase chain reaction using specific primers, and DNA sequencing was performed using the ABI 3710 Genetic Analyzer (Applied Biosystems, Foster City, CA).
3. Immunohistochemical analysisA single block of paraffin-embedded tissue from biopsy specimens was cut in serial 4-μm sections. For construction of tissue microarrays (TMAs) from surgically resected specimens, the most representative tumor areas were identified on a selected H&E slide and marked by a pathologist (J.Y.). Three replicate core samples, each measuring 2.0 mm in diameter, were obtained using a precise instrument, and arrayed on a recipient paraffin block. The first 4-μm sections were stained for H&E to verify histology.
Each case was tested using the following primary antibodies: DEL-specific monocloncal antibody (pre-diluted, clone SP111, Ventana Medical Systems Inc., Tucson, AZ), L858R-specific monoclonal antibody (pre-diluted, clone SP125, Ventana Medical Systems Inc.), and tEGFR antibody (1:100, clone SP9, Spring Bioscience, Pleasanton, CA). Briefly, 4-μm sections were deparaffinized in xylene, and rehydrated through a graded series of ethanol. Slides were labeled with antibody and protocol-specific bar codes, and loaded into a BenchMark XT automated slide stainer (Ventana Medical Systems Inc.). Incubation was performed for 16 minutes at 37°C with each primary antibody. The immunoreactions were detected using an Ultraview Universal DAB detection kit (Ventana Medical Systems Inc.) and 3,3′-diaminobenzidine, followed by counterstaining with hematoxylin and bluing reagent.
Immunohistochemical assessments were performed by an experienced pathologist (J.Y.) without information on molecular-based EGFR mutation status. Immunoreactivity was scored based on membranous and/or cytoplasmic staining, as follows: 0, no staining or faint staining in < 10% of tumor cells; 1+, weak staining in ≥ 10% of tumor cells; 2+, moderate staining in ≥ 10% of tumor cells; 3+, strong staining in ≥ 10% of tumor cells. For TMA samples, the staining scores obtained in three cores were averaged and the result was taken as a representative score for each case.
4. Statistical analysisIBM SPSS ver. 19.0 (IBM Co., Armonk, NY) was used for analysis. All variables were estimated using Pearson chi-square or Fisher exact tests. A p-value of < 0.05 was considered statistically significant. Immunohistochemical staining intensity (score 0-3) and predictive probability of the logistic regression model based on mutation-specific antibodies and tEGFR expression were used for construction of the receiver operating characteristic (ROC) curves and for calculation of the area under the curves (AUCs).
ResultsThe patient demographics and EGFR mutation status predefined by direct DNA sequencing are summarized in Table 1. There were 80 men and 74 women with a median age of 71 years (range, 37 to 89 years). Ninety-one patients (59.1%) were never-smokers, 12 (7.8%) were former smokers, and 51 (33.1%) were current smokers. The T status distribution was as follows: 18.8% T1, 34.4% T2, 38.3% T3, and 8.4% T4. None of the patients had received neoadjuvant chemotherapy or radiotherapy prior to the biopsy or tumor resection. Seventy-eight biopsies and 76 resected specimens were used in the current study. Direct DNA sequencing demonstrated EGFR mutations in 62 patients (40.3%). One patient (1.6%) had EGFR mutation in exon 18, 25 (40.3%) in exon 19, three (4.8%) in exon 20, and 33 (53.2%) in exon 21 (Table 2). The mutation detected in exon 18 was a double missense mutation. In exon 19, all were in-frame deletions: 14 (56%) with E746-A750 deletion and the remaining 11 (44%) with alternative in-frame deletions. In exon 20, one was of the duplication type and the others of missense type. In exon 21, L858R mutations were detected in 31 patients and alternative point mutations in three patients. E746-A750 deletion and L858R mutations accounted for 72.6% (45/62) of all EGFR mutations. Fig. 1 shows representative immunostaining with DEL-specific antibody, L858R-specific antibody, and tEGFR antibody. Immunoreactivity showed relatively homogeneous distribution between the cores of each tumor, and less than 9% had inconsistent findings in staining intensity with no effect on interpretation.
The AUCs for IHC with DEL-specific antibody and L858R-specific antibody were 0.843 with an optimal cutoff point of 0.5 (p=0.001) and 0.873 with an optimal cutoff point of 0.5 (p=0.001), respectively (data not shown). According to the optimal cutoff point of predictive probability by the logistic regression model, an immunostaining score of ≥ 1 was considered positive. The IHC data are shown in Tables 3 and 4. DEL-specific antibody correctly identified 10 of 14 patients with E746-A750 deletion. However, it was also positive in three patients with no E746-A750 deletion: one each with L745-A750 deletion in exon 19, L858R mutation, and EGFR wild type. There were four false-negatives for DEL-specific antibody. L858R-specific antibody correctly identified 24 of 31 patients with the corresponding gene mutation. Two EGFR wild type patients and one patient with T790M mutation in exon 20 were also positive for L858R-specific antibody, whereas seven patients with L858R mutation were negative for L858R-specific antibody. The overall sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) of DEL-specific antibody were 71.4%, 97.9%, 76.9%, and 97.2%, respectively (Table 5). L858R-specific antibody showed 77.4% sensitivity, 97.6% specificity, 88.9% PPV, and 94.5% NPV. No significant differences in mutant EGFR detection rate (23/78 [29.5%] vs. 22/76 [28.9%]; p > 0.05), or in sensitivity, specificity, PPV, and NPV were observed between biopsy samples and resected samples.
The sensitivity, specificity, PPV, and NPV of tEGFR antibody were 29%, 77.2%, 46.2%, and 61.7%, respectively (data not shown). No correlation was observed between tEGFR expression and EGFR mutation (p=0.284), DEL-specific antibody IHC (p=0.64), or L858R-specific antibody IHC (p=0.125) (Table 6).
DiscussionIn the current study, we compared the expression status of two EGFR mutation–specific antibodies between biopsies and resection samples from pulmonary adenocarcinoma, which had been previously genotyped by direct DNA sequencing. We also investigated tEGFR protein for evaluation of its possible correlation with mutant EGFR proteins detected by mutation-specific antibodies. The major findings of our study are as follows: (1) IHC with mutation-specific antibodies correctly identified EGFR mutation status in 89% (137/154), showing negative immunostaining in 94.5% (103/109) of patients with EGFR wild type or mutations other than E746-A750 deletion and L858R mutation, and positive reaction in 75.6% (34/45) of patients with corresponding mutations; (2) no significant differences in the EGFR mutation frequency were observed between biopsies and resection specimens; (3) tEGFR expression showed no association with immunoreactivity for both mutation-specific antibodies.
Yu et al. [3] developed a set of EGFR mutation–specific antibodies against EGFR with E746-A750 deletion and L858R for identification of mutant EGFR proteins by IHC, and reported a sensitivity of 92% and specificity of 99% as compared with direct DNA sequencing. Although accumulating data have suggested IHC using the two mutation-specific monoclonal antibodies as a simple, rapid, and costeffective screening tool for discriminating lung cancer patients responsive to EGFR-based therapies [4-10], the reported sensitivities and specificities were quite variable. In several investigations on NSCLC, the specificities of both antibodies were relatively high (77%-100%), whereas the sensitivities of DEL-specific and L858R-specific antibodies ranged from 40% to 94% and 24% to 100%, respectively [5,6,9,10,15-17]. All used the same two antibodies (clone 43B2 specific for E746-A750 deletion and clone 6B6 specific for L858R mutation, Cell Signaling Technology, Danvers, MA), so that the wide range of sensitivities and specificities is most likely not due to the antibody used. In addition, we used EGFR mutation-specific antibodies purchased from a different manufacturer (clones SP111 and SP125, respectively, Ventana Medical Systems Inc.). The overall sensitivity was 75.6% with a specificity of 94.5%. DEL-specific IHC showed a sensitivity of 71.4% and specificity of 97.1%, while L858R-specific IHC showed a sensitivity of 77.4% and specificity of 97.6%, compatible with previous reports. Similar results were obtained by the most recent investigation using the same antibodies as in the current study [18]. The wide range of sensitivities and specificities may be attributed to scoring systems, which play a critical role in IHC-based analyses. Several scoring schemes for interpretation of mutation-specific IHC have been adopted in published papers. Immunoreactivity was classified on the basis of staining intensity, sometimes multiplied by the percentage values, both of which are highly dependent on the researcher. Another matter is the distribution of staining: membrane or cytoplasm. Some assessed immunostaining in membrane only, others in membrane and/or cytoplasm, or even based on cytoplasmic staining of tumor cells alone [3,7,9,15,19]. Furthermore, when using a four-grading scale (score 0-3), ≥ 1+ was defined as positive by some investigators, and ≥ 2+ was considered positive by others [7-10,19]. Recently, an optimized protocol of an intensity of 2+ or more in membrane and/or cytoplasm of > 10% tumor cells for positivity was shown to be the most appropriate way to interpret the EGFR mutation–specific IHC [10]. In our study, we performed ROC analyses to determine the best cutoff for detection of EGFR mutations, and found it to be a score of 1+ or more. Consensus of universally accepted criteria in interpreting the results as positive or negative should be reached in order to produce objective and reproducible results before using mutation-specific IHC as a substitute for molecular methods.
There have been conflicting results regarding the immunostaining pattern of mutation-specific antibodies. Kitamura et al. [9] observed different staining intensity in at least one of three TMA cores each from 15/33 positive tumors (45%). On the other hand, Xiong et al. [10] used whole tissue sections and reported homogeneity in staining pattern. This prompted us to investigate whether the type of specimen (biopsy vs. resection) would make a difference in determining mutation status by IHC. In the current study, observed staining patterns were homogeneous in both biopsies and TMA cores obtained from each patient. In addition, no significant differences in mutant EGFR protein detection rates were found between the biopsy samples and TMAs (29.5% vs. 28.9%, p > 0.05). Fan et al. [11] described a heterogeneous expression pattern in some of their cases with no details given, and speculated the causal relationship between heterogeneity and lower sensitivity of DEL-specific antibody in their biopsy samples. However, they also detected the EGFR mutation phenotype in 32.1% of biopsies and 36.3% of resection samples (p > 0.05) with high specificity (97.9%-100%), which is in concordance with our results, suggesting that IHC using two mutation-specific antibodies may be performed as reliably in biopsies as in resection samples. Therefore, when biopsied specimens are the only materials available for EGFR status in patients at advanced stage, and tumor cells in the samples are insufficient or inadequate for DNA-based molecular methods, using IHC with mutation-specific antibodies may preclude another biopsy procedure to obtain ample tissues.
The overall sensitivity of IHC with DEL-specific antibody for all exon 19 mutations was low (44%). This is because DEL-specific antibody is relatively specific for the frequent E746-A750 deletion and cannot detect the alteration [3,8]. In the current study, 11 of 25 patients with exon 19 mutations had alternative in-frame deletions other than E746-A750 deletion, and thus almost exclusively did not react with DEL-specific antibody. DNA sequencing analyses should always be performed if IHC shows negative results.
Our observation of the lack of correlation between EGFR mutation and tEGFR IHC is not surprising. Identification of tEGFR protein by IHC did not always reflect the detection of EGFR mutation, suggesting that mechanisms other than EGFR mutation may also play a role in tEGFR expression. As in the study herein, all previous publications, to the best of our knowledge, used tEGFR antibodies detecting the external domain (ED) of EGFR. We found no correlation between tEGFR and mutation-specific antibody expression. Kitamura et al. [9] reported a correlation of tEGFR with mutation-specific antibodies, using another ED-specific antibody (clone 2-18C9, DAKO, Carpinteria, CA) for tEGFR. However, they included histologic types other than adenocarcinoma. A histologically inhomogeneous series of lung cancer may be one possible explanation for this discrepancy. ED-specific antibodies bind to receptors containing ED and do not discriminate between active and inactive receptors, whereas intracellular domain (ID)–specific antibody would detect active or even truncated forms of the receptor. One study demonstrated that EGFR expression evaluated by ED-specific antibody did not predict patient survival, but EGFR protein expression using an ID-specific antibody was a significant predictor of response to gefitinib in NSCLC patients [20]. Investigations on whether EGFR ID-specific antibody IHC is correlated with mutant EGFR, and has an impact on diagnostic power in the IHC interpretation of the mutation-specific antibodies, may be of great help in more accurate assessment of EGFR mutation status. Such studies are currently in progress at our laboratories.
ConclusionIHC with EGFR mutation–specific antibodies exhibited extremely high specificity (94.5%) with good sensitivity (75.6%) in both biopsies and resection specimens. Therefore, IHC using EGFR mutation–specific antibodies may preclude a second biopsy procedure to obtain additional tissues for identification of EGFR mutations by molecular biology techniques in biopsies from advanced cancer, especially when tumor cells in the samples are limited. Negative cases, however, require further molecular-based analyses for identification of other mutations with low frequency.
Fig. 1.Representative immunostaining photomicrographs of a tumor with epidermal growth factor receptor (EGFR) wild type (A-C), a tumor with E746-A750 deletion (DEL) (D-F), and a tumor with L858R mutation (G-I) detected by DNA sequencing, stained with total EGFR antibody (A, D, G), with DEL-specific antibody (B, E, H), and with L858R-specific antibody (C, F, I). Positive immunoreactivity with membrane reinforcement is observed in panels D, E, and I (A-I, ×200). 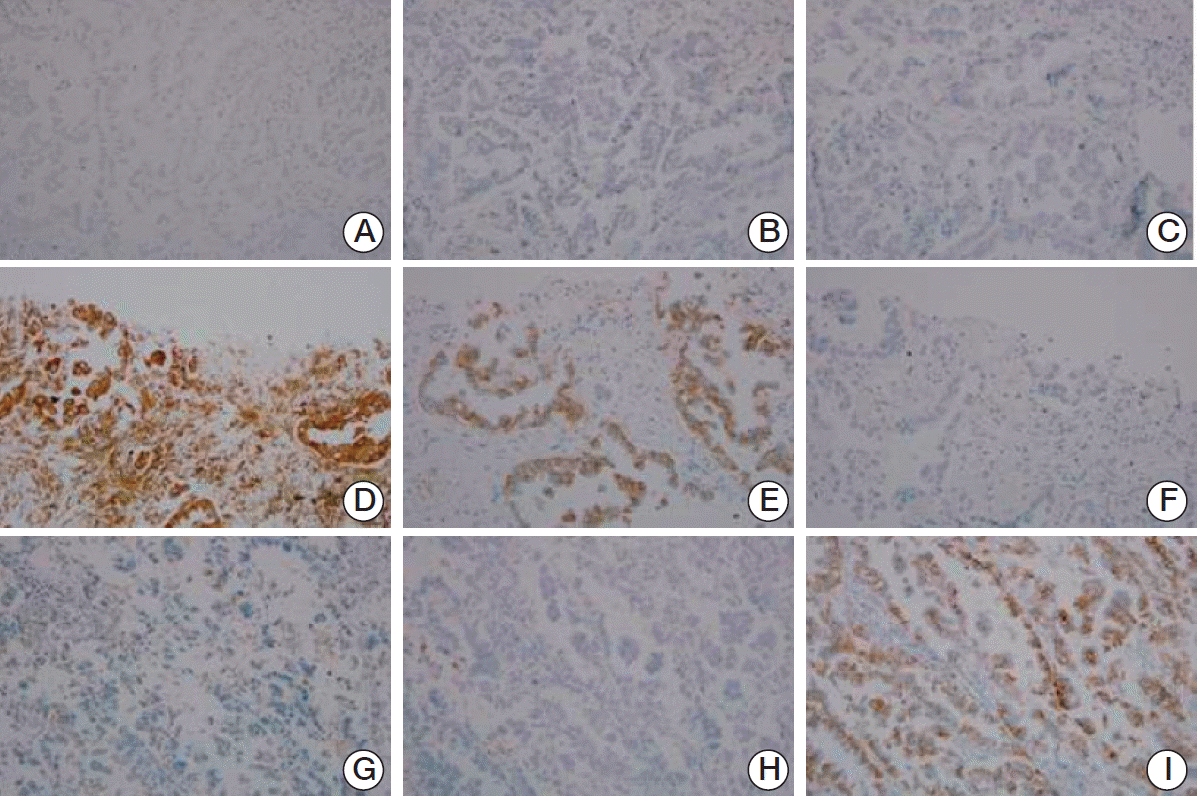
Table 1.Clinicopathologic characteristics and EGFR DNA sequencing results (n=154) Table 2.
EGFR mutation types detected in 62 patients Table 3.Comparison of immunohistochemistry and direct DNA sequencing analysis for EGFR mutation Table 4.List of discordant cases between immunohistochemistry and direct DNA sequencing Table 5.Detection accuracy of EGFR mutation–specific immunohistochemistry Table 6.Comparison of immunostaining results using three different antibodies References1. Riely GJ, Pao W, Pham D, Li AR, Rizvi N, Venkatraman ES, et al. Clinical course of patients with non-small cell lung cancer and epidermal growth factor receptor exon 19 and exon 21 mutations treated with gefitinib or erlotinib. Clin Cancer Res. 2006;12(3 Pt 1):839–44.
2. Mok TS, Wu YL, Thongprasert S, Yang CH, Chu DT, Saijo N, et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N Engl J Med. 2009;361:947–57.
3. Yu J, Kane S, Wu J, Benedettini E, Li D, Reeves C, et al. Mutation-specific antibodies for the detection of EGFR mutations in non-small-cell lung cancer. Clin Cancer Res. 2009;15:3023–8.
4. Kawahara A, Taira T, Azuma K, Tominaga M, Hattori S, Kawahara M, et al. A diagnostic algorithm using EGFR mutation-specific antibodies for rapid response EGFR-TKI treatment in patients with non-small cell lung cancer. Lung Cancer. 2012;78:39–44.
5. Kato Y, Peled N, Wynes MW, Yoshida K, Pardo M, Mascaux C, et al. Novel epidermal growth factor receptor mutation-specific antibodies for non-small cell lung cancer: immunohistochemistry as a possible screening method for epidermal growth factor receptor mutations. J Thorac Oncol. 2010;5:1551–8.
6. Simonetti S, Molina MA, Queralt C, de Aguirre I, Mayo C, Bertran-Alamillo J, et al. Detection of EGFR mutations with mutation-specific antibodies in stage IV non-small-cell lung cancer. J Transl Med. 2010;8:135.
7. Ambrosini-Spaltro A, Campanini N, Bortesi B, Azzoni C, Naldi N, Ampollini L, et al. EGFR mutation-specific antibodies in pulmonary adenocarcinoma: a comparison with DNA direct sequencing. Appl Immunohistochem Mol Morphol. 2012;20:356–62.
8. Kawahara A, Azuma K, Sumi A, Taira T, Nakashima K, Aikawa E, et al. Identification of non-small-cell lung cancer with activating EGFR mutations in malignant effusion and cerebrospinal fluid: rapid and sensitive detection of exon 19 deletion E746-A750 and exon 21 L858R mutation by immunocytochemistry. Lung Cancer. 2011;74:35–40.
9. Kitamura A, Hosoda W, Sasaki E, Mitsudomi T, Yatabe Y. Immunohistochemical detection of EGFR mutation using mutation-specific antibodies in lung cancer. Clin Cancer Res. 2010;16:3349–55.
10. Xiong Y, Bai Y, Leong N, Laughlin TS, Rothberg PG, Xu H, et al. Immunohistochemical detection of mutations in the epidermal growth factor receptor gene in lung adenocarcinomas using mutation-specific antibodies. Diagn Pathol. 2013;8:27.
11. Fan X, Liu B, Xu H, Yu B, Shi S, Zhang J, et al. Immunostaining with EGFR mutation-specific antibodies: a reliable screening method for lung adenocarcinomas harboring EGFR mutation in biopsy and resection samples. Hum Pathol. 2013;44:1499–507.
12. Dimou A, Agarwal S, Anagnostou V, Viray H, Christensen S, Gould Rothberg B, et al. Standardization of epidermal growth factor receptor (EGFR) measurement by quantitative immunofluorescence and impact on antibody-based mutation detection in non-small cell lung cancer. Am J Pathol. 2011;179:580–9.
13. Selvaggi G, Novello S, Torri V, Leonardo E, De Giuli P, Borasio P, et al. Epidermal growth factor receptor overexpression correlates with a poor prognosis in completely resected non-small-cell lung cancer. Ann Oncol. 2004;15:28–32.
14. Ceppi P, Volante M, Novello S, Rapa I, Danenberg KD, Danenberg PV, et al. ERCC1 and RRM1 gene expressions but not EGFR are predictive of shorter survival in advanced non-small-cell lung cancer treated with cisplatin and gemcitabine. Ann Oncol. 2006;17:1818–25.
15. Kozu Y, Tsuta K, Kohno T, Sekine I, Yoshida A, Watanabe S, et al. The usefulness of mutation-specific antibodies in detecting epidermal growth factor receptor mutations and in predicting response to tyrosine kinase inhibitor therapy in lung adenocarcinoma. Lung Cancer. 2011;73:45–50.
16. Brevet M, Arcila M, Ladanyi M. Assessment of EGFR mutation status in lung adenocarcinoma by immunohistochemistry using antibodies specific to the two major forms of mutant EGFR. J Mol Diagn. 2010;12:169–76.
17. Hofman P, Ilie M, Hofman V, Roux S, Valent A, Bernheim A, et al. Immunohistochemistry to identify EGFR mutations or ALK rearrangements in patients with lung adenocarcinoma. Ann Oncol. 2012;23:1738–43.
18. Seo AN, Park TI, Jin Y, Sun PL, Kim H, Chang H, et al. Novel EGFR mutation-specific antibodies for lung adenocarcinoma: highly specific but not sensitive detection of an E746_A750 deletion in exon 19 and an L858R mutation in exon 21 by immunohistochemistry. Lung Cancer. 2014;83:316–23.
|
|
|||||||||||||||||||||||||||||||||||||||||||