Introduction
Molecular classification plays a very important role in decision-making regarding treatment of colorectal cancer and in prediction of the prognosis. However, identification of these mutations is usually possible by postoperative testing of tissues removed at the time of surgery. Therefore, the opportunity to confirm the presence of genetic mutations preoperatively is limited. Chemotherapy and radiotherapy are widely used preoperatively in the treatment of rectal cancer, and sometimes no tumor cells are found in surgical tissue specimens. Therefore, it would be helpful to be able to confirm genetic mutation on imaging without the need for histopathologic examination when making decisions regarding treatment. The Kirsten Ras (KRAS) gene mutation is one of several common mutations in colorectal cancer and is found in 30%-50% of cases. Several studies have indicated that the presence of mutant KRAS in colorectal cancer correlates with a poor response to epidermal growth factor receptor inhibitors in a metastatic setting [1,2].
Radiomics involves high-throughput extraction of large amounts of imaging features from radiologic images and has recently emerged as a promising tool for predicting the prognosis and guidance of therapy [3-5]. Moreover, imaging features identified by magnetic resonance (MR)–based radiomics analysis were demonstrated to effectively predict the response of rectal cancer to chemoradiotherapy [4,6-10]. Radiogenomics is an emerging field that integrates imaging and genomics data to identify imaging correlates of a specific tumor genotype or molecular phenotype for precision medicine [5]. Several studies have assessed the associations of positron emission tomography (PET)–based or computed tomography (CT)–based texture features with KRAS mutations in colorectal cancer and rectal cancer [11-13]. To the best of our knowledge, there have been no studies on whether or not the T2-weighted MR-based radiomics signature is specifically associated with KRAS mutation status in rectal cancer. In recent years, biomarkers detected on MR imaging have shown potential for identifying tumor characteristics and improving our ability to diagnose certain tumors, including rectal cancer [14]. The aim of this study was to investigate the potential of T2-weighted MR imaging features to identify KRAS mutation status in rectal cancer.
Materials and Methods
1. Study population
The study population initially comprised samples from 82 patients that were collected retrospectively from the tumor bank for a molecular genomics study between December 2008 and January 2010 (n=52) or collected prospectively from patients with mid to lower rectal cancer for a radiogenomics study between September 2017 and March 2018 (n=30). Twenty-two of the 82 patients were subsequently excluded, leaving data for 60 patients for inclusion in the analyses. The study inclusion and exclusion criteria and the flow of patients through the study are shown in Fig. 1.
The mean patient age was 61.2 years and the mean body mass index was 24.2. An MR imaging examination was performed in all cases before treatment for rectal cancer was started. Eighteen patients (30%) received preoperative chemoradiation. Samples collected retrospectively from the tumor bank were obtained at the time of surgery whereas the prospectively collected tissue samples were obtained by endoscopic biopsy using large-cup jumbo forceps before starting treatment in patients who received preoperative chemoradiation. Two of 24 tissue samples obtained by endoscopic biopsy did not meet the quality control standard for genetic testing (Fig. 1). The demographic and clinical characteristics of the 60 patients are shown in Table 1.
2. DNA test
DNA for detection of KRAS mutation status was isolated from fresh frozen tissue using an Automated Nucleic Acid Extractor (Roche Molecular Biochemicals, Mannheim, Germany). The purified genomic DNA from the samples was amplified by polymerase chain reaction. MassArray with matrix-assisted laser desorption/ionization-time of flight mass spectrometry and an iPLEX ADME PGx panel (Agena Bioscience, San Diego, CA) modified by the Omics Core Laboratory at the National Cancer Center were used for detection of mutations. Eleven mutations in the KRAS gene (A59T, G12A, G12C, G12D, G12F, G12R, G12S, G12V, G13D, G61H, and Q61L) were detected in this panel. Twenty-five patients were found to have mutant KRAS and 35 to have wild-type KRAS.
3. MR imaging
MR imaging was performed using a 3-T scanner (Achieva 3.0T, Philips Healthcare, Amsterdam, The Netherlands; n=39; Achieva Tx 3.0T, Philips Healthcare, Best, The Netherlands; n=13; Ingenia Cx 3.0T, Philips Healthcare, Best, The Netherlands; n=8) and a 32-channel phased array body coil (USA Instruments Inc., Aurora, OH) using the standard imaging protocol. One hour before the MR imaging examination, one bisacodyl suppository (Dulcolax, Boehringer Ingelheim, Ingelheim am Rhein, Germany) was administered for bowel preparation. Thirty minutes before MR imaging, 20 mg of scopolamine butylbromide (Buscopan, Boehringer Ingelheim) was injected intravenously to reduce colonic motility, unless contraindicated. Standard T2-weighted, fast spin-echo MR sequences were obtained in the sagittal, axial, and coronal planes using the following parameters: echo time/repetition time, 80-110/2,500-8,600 msec; slice thickness, 3-5 mm; echo train length, 16-32; matrix, 224×224 to 800×538; and field of view, 150×150 to 360×360. Axial, T1-weighted, three-dimensional spoiled-gradient-echo sequences and diffusion-weighted images were also obtained.
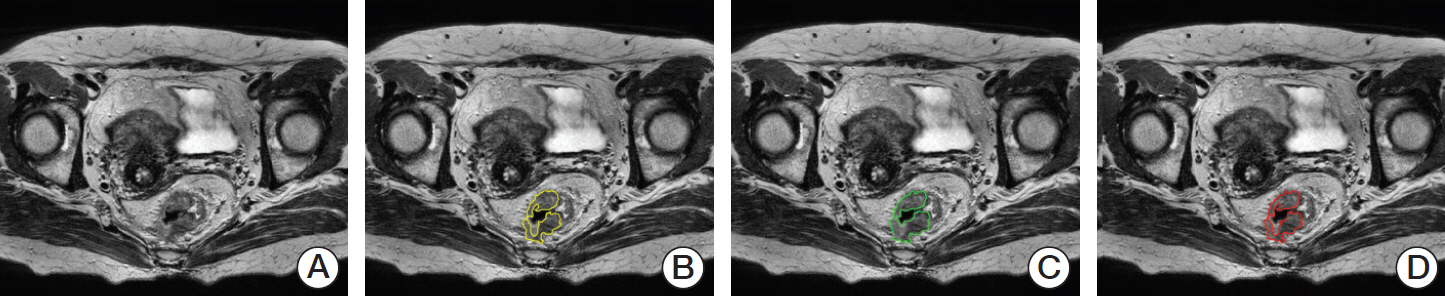
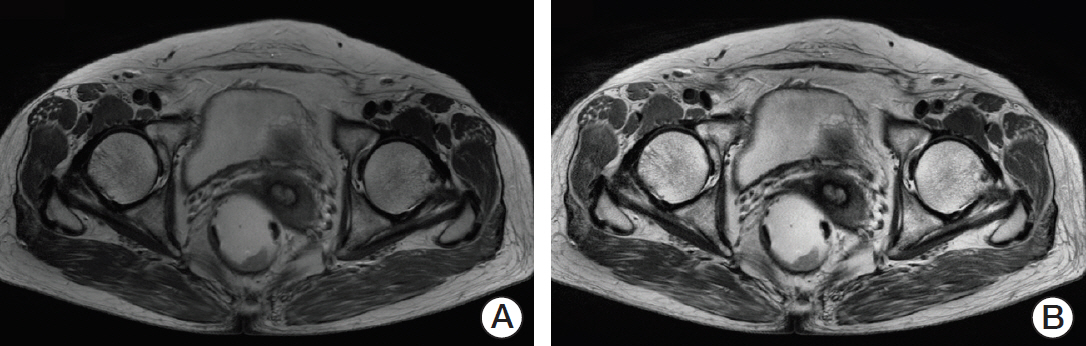
We selected the slice with the largest cross-sectional area of the tumor as the representative slice. A region of interest (ROI) indicating the tumor boundaries was manually drawn by two experienced radiologists on each representative slice, as shown in Fig. 2. On T2-weighted MR images, tumors were defined as areas that were isointense to hyperintense in comparison with the relatively hypointense adjacent normal muscular rectal wall. Texture analysis was performed on all ROIs (manually segmented by two experienced radiologists) in T2-weighted MR images for 60 patients. The ground truth tumor region was defined by the overlapping areas of two ROIs that were drawn independently by two radiologists, as shown in Fig. 2. If there was uncertainty regarding the region of the tumor, the area was not included in the segmentation.
4. Image preprocessing
Preprocessing of the proposed algorithm can be divided into two major steps, i.e., normalization of the image intensity range and histogram equalization (Fig. 3). To normalize the image intensity range, all images were converted from 12-bit to 8-bit grayscale using the maximum and minimum pixel value over the body region of each patient. Contrastlimited adaptive histogram equalization was also applied to enhance the contrast in the rectum and reduce the effect of magnetic bias [15-17].
5. Texture analysis
For the quantitative analysis, we extracted five groups of texture features with tumor ROIs using the MATLAB program (MathWorks Inc., Natick, MA). The five groups consisted of histogram (n=4), gradient (n=4), gray-level co-occurrence matrix (GLCM; n=5), gray-level run length matrix (n=7), and Laplacian of Gaussian (LoG) filtered features (n=4 for each filter size). The most common histogram features are simply the statistical parameters of the histogram distribution and consist of the mean (average brightness), standard deviation (SD), kurtosis (a measure of peakedness and tailedness), and skewness (a measure of asymmetry of the histogram). The image gradient feature of the histogram is computed for the 3×3-pixel neighborhood and the parameters of such a histogram distribution are determined [18,19]. These gradient features are Gmean, G_SD, Gkurtosis, and Gskewness.
The LoG band-pass filtration technique enhanced and extracted features of different sizes based on a spatial scale filter (SSF) value, which varies from fine texture (SSF1 and SSF2, 1 mm and 2 mm in radius, respectively), to medium texture (SSF3 and SSF4, 3 mm and 4 mm), to coarse texture (SSF5 and SSF6, 5 mm and 6 mm). Next, the filtered texture maps were quantified using the histogram parameters, i.e., the mean, kurtosis, skewness, and SD [20,21]. These histogram parameters were also quantified using the conventional MR image without filtration (i.e., SSF0).
GLCM is an efficient texture analysis method that uses second-order statistics to characterize two adjacent pixel values at specific locations. The GLCM features used in this study were correlation, angular second moment, homogeneity, and entropy [22-24].
A gray-level run length matrix (GLRLM) is a texture representation model that extracts the spatial plane features of each pixel. Each element in this matrix gives the total number of occurrences of the gray level in a given direction. In this study, we selected seven GLRLM features, i.e., short run emphasis, long run emphasis, gray-level nonuniformity, run length nonuniformity, run percentage, low gray-level run emphasis, and high gray-level run emphasis [25-28].
6. Statistical analysis
We selected the prospective features from T2-weighted MR images using the two-tailed t test method. The features selected were used to build a discriminant model using the decision tree method to estimate whether the KRAS mutation was present. The decision tree was developed using the classification and regression tree method, which is an empirical, statistical technique based on recursive partitioning analysis [29] and generates binary decision trees. Using the decision tree method, diagnostic threshold values were determined iteratively to maximize the diagnostic accuracy while minimizing false-positive KRAS mutations. The tree does not expand its node unless the Gini index does not improve by more than 0.001. To restrict overfitting, minimum numbers of cases for nodes are set to 20 for the parent node and five for the child node. Therefore, no further separations of the tree were used if an output arm of the decision node contained five or fewer patients within either diagnostic category, i.e., mutant KRAS and wild-type KRAS. We evaluated the performance of the final decision tree by receiver-operating characteristic curve analysis and the sensitivity, specificity, and accuracy values. The sensitivity value indicates the ability of the decision tree to identify the KRAS mutation correctly. All statistical analyses were performed using SPSS software ver. 14.0 (SPSS Inc., Chicago, IL). A p-value of < 0.05 was considered statistically significant.
Results
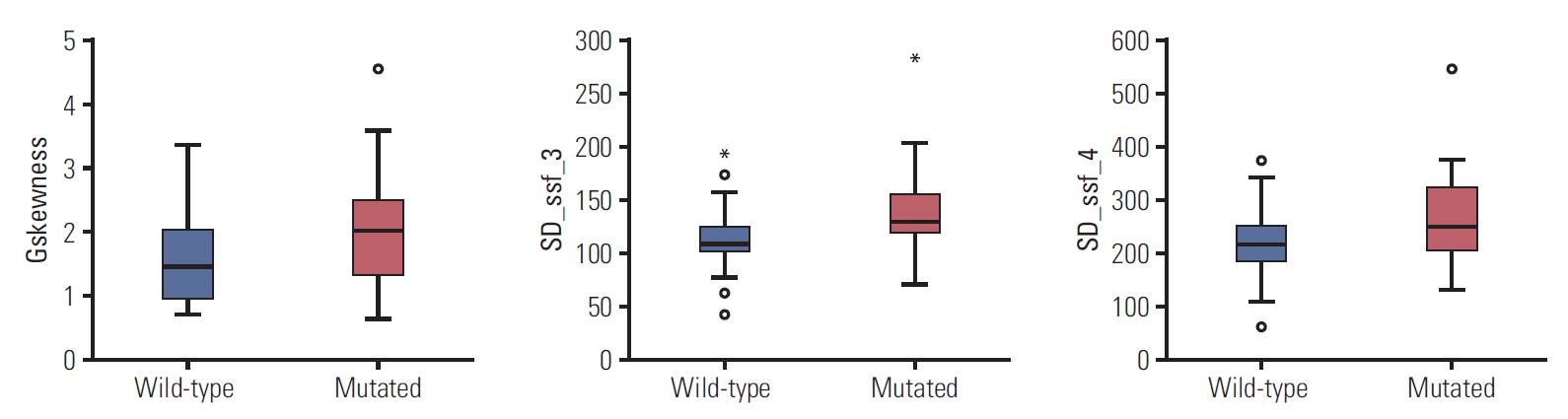
We extracted the texture features from the T2-weighted MR images (S1 Table). Only three radiomics features were significantly associated with KRAS mutation status (p < 0.05). The selected imaging features are summarized in Table 2. This table also compares the distribution of the three imaging features according to KRAS mutation status. There was a statistically significant difference in three imaging features between patients with mutant KRAS and those with wild-type KRAS (p < 0.05). The mean (and SD) value for skewness with gradient filter (Gskewness) was significantly higher in the KRAS mutation group than in the wild-type group (2.04±0.94 vs. 1.59±0.69). A higher SD at medium texture (SSF3 and SSF4) had a significant ability to differentiate mutant KRAS (139.81± 44.19 and 267.12±89.75, respectively) from wild-type KRAS (114.55±29.30 and 224.78±62.20). The box plot in Fig. 4 shows the differences in the three imaging features.
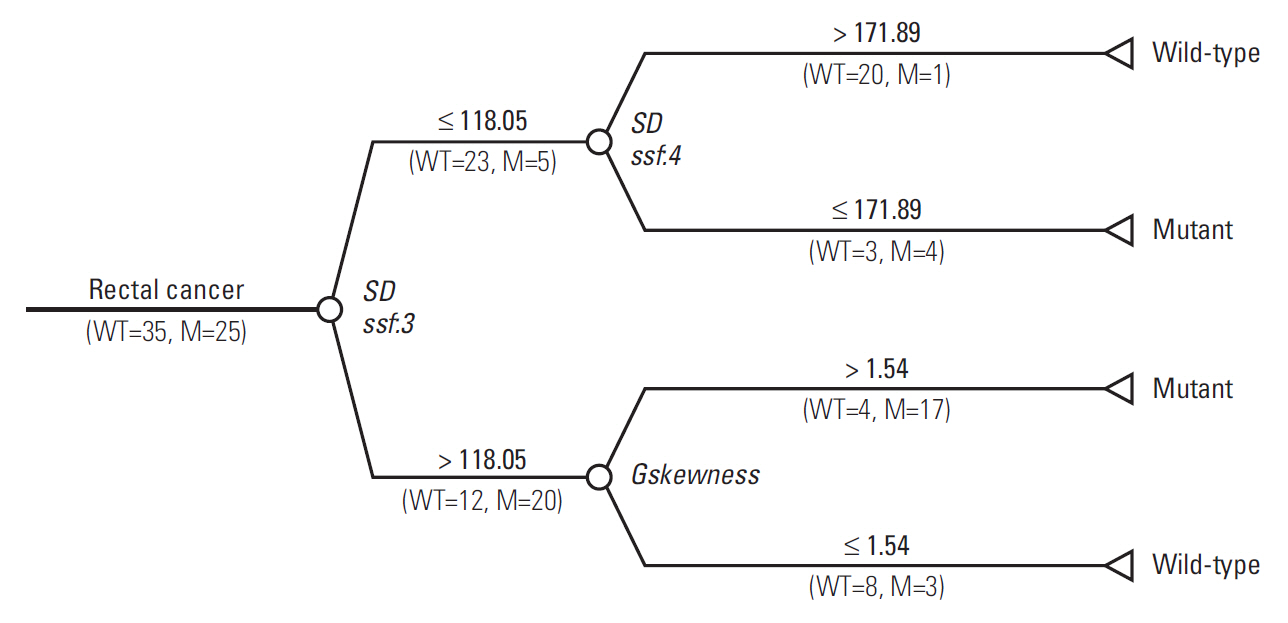
The final decision tree comprised four nodes as shown in Fig. 5. The first node of the decision tree used the SD at medium texture with a threshold value of 118.05. The SD at medium texture was assessed second and the Gskewness of the gradient feature was assessed last. Therefore, the final decision tree comprised three decision nodes and four terminal nodes (of which two identified KRAS mutation and the remaining two identified wild-type status). The sensitivity, specificity, and accuracy values of the decision tree for the whole dataset were 84%, 80%, and 81.7%, respectively.
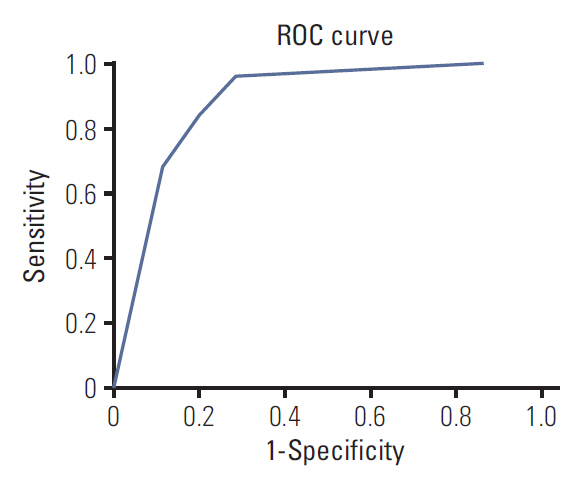
The receiver-operating characteristic curves for the decision tree model are shown in Fig. 6. The area under the curve was 0.884.
Discussion
In this study, we used a decision tree model for identification of KRAS mutations using T2-weighted MR imaging features to decide on the treatment method and predict the prognosis of rectal cancer. In general, invasive biopsy or surgical procedures are needed to identify the mutation status of KRAS. T2-weighted MR imaging-based texture analysis could be used as an alternative noninvasive strategy to overcome these limitations.
Several studies have attempted to investigate the relationship between imaging characteristics and KRAS mutation. Miles et al. [11] showed that a combination of PET and CT imaging features can potentially provide an imaging signature for KRAS mutation in colorectal cancer. In that study, the true-positive rate, false-positive rate, and accuracy of the proposed decision tree model was 82.4%, 0%, and 90.1%, respectively [11]. Yang et al. [12] found significant associations between a CT-based radiomics signature and KRAS/NRAS/BRAF mutations. The sensitivity, specificity, and area under the curve for prediction of KRAS/NRAS/BRAF mutations were 0.757, 0.833, and 0.869, respectively, in their primary cohort. However, there was no association of background clinical characteristics, tumor stage, or histological differentiation with KRAS/NRAS/BRAF mutation status. Kawada et al. [13] found that 18F-fluorodeoxyglucose PET/CT was useful for prediction of KRAS status in metastatic colorectal cancer with a sensitivity, specificity, and accuracy of 68%, 74%, and 71.4%, respectively. Meng et al. [30] showed that the radio-mics signatures based on multiparametric MR imaging predicted the KRAS gene mutation with an area under the curve (AUC) of 0.651. Cui et al. [31] found that diffusion kurtosis imaging-derived histogram metrics from whole tumor volumes were associated with KRAS mutations. Most of the K parameters showed moderate diagnostic significance for KRAS mutations. The 75th percentile pixel value of K showed the highest AUC of 0.871 and the sensitivity, specificity, positive predictive value, and negative predictive value were 81.43%, 78.21%, 77.03%, and 82.43%, respectively [31]. Similarly, Xu et al. [32] showed that diffusion-weighted MR imaging-derived para-meters can predict KRAS mutation in rectal cancer. They found that the mean apparent diffusion coefficient and D* (pseudo-diffusion coefficient) had moderate diagnostic significance, with respective AUCs of 0.756 and 0.710 [32]. Yeo et al. [33] reported that the perfusion parameters from dyna-mic contrast-enhanced MR imaging were significantly associated with KRAS mutation. The mean and mode Ktrans parameters demonstrated high diagnostic performance for KRAS gene mutation with respective AUCs of 0.788 and 0.793 [33]. To our knowledge, this is the first report on the relationship between T2-weighted MR-based texture features and KRAS mutation. MR imaging can reduce the impact of image noise, which can affect texture analysis because of the higher contrast resolution and contrast-to-noise ratio when compared with CT [7]. However, previous studies have shown that pretreatment T2-weighted MR imaging radiomics has the potential to characterize the histopathologic features of rectal cancer and predict the response to chemoradiotherapy, except for KRAS mutation [4,9,14,34]. Therefore, in this preliminary study, we investigated the potential association between T2-weighted MR imaging features and KRAS mutation status.
Several studies have suggested that texture features may be related to the tumor microenvironment and the presence of hypoxia or angiogenesis [11,35-37]. A hypoxic status leads to an increase in the aggressiveness of the tumor and resistance to treatment [38]. Ganeshan et al. [36] found greater spread in hypoxic tissues and more variation in voxel values on contrast-enhanced CT. We found that KRAS mutations were associated with higher Gskewness and larger SD values for medium texture. Gskewness and SDs are associated with variation in intensity. Consequently, higher Gskewness and larger SD values indicate that the group with KRAS mutation is more heterogeneous than the group with wild-type KRAS. For the prediction of KRAS mutation status, we used a decision tree model that can deal with large, complicated datasets efficiently without imposing a complicated parametric structure. This decision tree is also easy to understand and implement. Using decision tree analysis based on these imaging features, we were able to identify MR-based imaging features that identified KRAS mutations. The sensitivity, specificity, and accuracy of the decision tree for differentiating tumors with mutant KRAS from those with wild-type KRAS were 84%, 80%, and 81.7%, respectively. The model used in the present study has better performance in predicting KRAS mutation status than that used in previous research [11-13].
This study has several limitations. First, the proposed model needs to be externally validated by a prospective study in the future using patients whose data are not used to construct the model. Second, the number of cases included in this study was relatively small, so the reliability of the proposed model needs to be determined in a future study that includes more cases. Third, radiomics was performed only on T2-weighted MR images. Radiomics analysis may be improved with inclusion of other MR imaging sequences, such as diffusion-weighted imaging and dynamic contrast-enhanced MR imaging. In this study, KRAS mutation in rectal cancer was confirmed on MR images. Arguably, this study is of limited value because it is relatively easy to perform endoscopic biopsy in rectal cancer. However, our findings indicate that post-histologic examination is helpful in confirming serial gene mutation changes during treatment for rectal cancer.
In conclusion, our study shows that T2-weighted MR imaging has potential as a noninvasive imaging modality to predict KRAS mutation status in patients with rectal cancer before surgery. The proposed model may also help to determine the best treatment strategy for individual patients with rectal cancer.